April 21, 2022
I. introduction
It is a popular misconception that the human species has stopped evolving. At the other extreme, some colorful speculative fiction has imagined radically evolved future humans with supernatural powers, dramatic physiological changes, some robotic features, or even branching into two or more very distinct descendant species. In this post we will review what science tells us about the past evolution of homo sapiens following their migration out of Africa some 50,000 to 100,000 years ago, and even since the establishment of the first human civilizations some 6,000 years ago, as well as what speculations about future human evolution modern science supports.
Our knowledge of past human evolution has advanced dramatically this century with our ability to perform DNA sequencing not only on modern humans, but also on the skeletal remains of our ancestors and of the closely related hominin species of Neanderthals and Denisovans. Natural evolution is generally a very slow process, although we will point out in this post a couple of circumstances when it may proceed faster. Many tens or even hundreds of generations are typically required before one can detect clear evolutionary changes in a species. In this context, it is worth noting that the entirety of human civilizations to date has spanned only several hundred human generations.
Thus, the past evolutionary changes we can detect in homo sapiens reflect evolutionary pressures experienced in ancient human communities. There has been insufficient time to date for natural evolution to respond to more recent, post-industrial changes in human conditions, requiring adaptation to dense urban populations, new diseases and dietary trends, different social, cultural, environmental, and demographic pressures, and impacts of modern technology. We can make educated guesses of future human evolution based on natural evolutionary trends needed for optimal adaptation to these recent societal developments. But the natural processes we will outline in section II are now supplemented by new technologies that add “human selection” as an important new evolutionary pressure, whose future impacts are more challenging to predict because the technology is advancing so rapidly.
We rely heavily in our discussions on two recent books covering these topics: Future Humans: Inside the Science of our Continuing Evolution by evolutionary biologist Scott Solomon and Future Humans: What Will Humans Look Like in a Million Years? by Professor of bioinformatics Thomas Mailund. Our post has two parts: in the first, we will discuss the various processes that contribute to evolutionary change (section II) and evidence of past human evolution (section III); in the second part, we will give examples of how humans may evolve in the future through natural evolutionary processes (section IV) and through applications of contemporary high technology (section V), before summarizing in section VI.
II. evolutionary processes
Any significant changes to the human genome, averaged over individuals either globally or in specific locations, can be considered as an evolutionary change. The human genome comprises 23 chromosomes encompassing some 3 billion pairs of ribonucleotide bases, including 20,000 or so genes that influence phenotype – i.e., structure, appearance and behavior — and encode the assembly of proteins and regulatory processes necessary for life functions. While scientists have mapped the entire human genome, we still have an incomplete understanding of which genes contribute to which phenotypic features. Each individual has two genomes, one contributed by each biological parent. Each gene can come in a number of variants, called alleles. Alleles may differ from one another by changes to a single base pair (called single-nucleotide polymorphisms, SNP) or multiple base pairs.
The vast majority of possible homo sapiens alleles already exist at some level within today’s global human population of some 7.9 billion people. Evolution occurs when the frequency of occurrence of different alleles within the human population undergoes substantial changes over time. As we explain below and have covered to some extent in our previous post on evolution, some such changes are naturally or sexually selected to enhance fitness, while others may occur randomly, or through mating or sudden loss of different population segments, or nowadays through genetic engineering to implement human preferences.
In the evolutionary context, fitness is measured by a species’ success in producing descendants. Following an example used by Mailund, consider a population that initially consists of two equal size groups: group A, whose females consistently produce three children reaching child-bearing age, and group B, whose females produce two such children. The next generation will see group A representing 60% of the population. That fraction will rise in the following generation to 69%, and by the 10th generation to more than 98% of the resulting (now much larger than initial) population. The bottom line of evolution is this: the most fertile dominate future gene pools, so long as their young live long enough to produce their own offspring.
Mutation and natural selection:
When living cells with DNA-based chromosomes divide, each daughter cell has to be supplied with a copy of the parent cell’s DNA. The replication process is the basis for biological inheritance, but the fidelity of replication is not perfect. The error rate in human self-correcting DNA replication has been estimated to be about one per ten billion base pairs, which sounds awfully good. But the full human genome, summed over all 23 chromosomes, comprises about three billion base pairs. Thus, there is a roughly 30% probability of getting at least one mutation per genome per cell division. The generation of human sperm generally involves hundreds of cell divisions. The net result is that each new human born is estimated to have, on average, about 70 new DNA mutations compared to the parents. The most common type of mutation is a change to a single base pair (SNP), but others involve sections of DNA which get moved, inserted, deleted or inverted during replication.
In a global human population of nearly 8 billion people, there can thus be some 560 billion mutations relative to the preceding generation. Since the human genome comprises about 3 billion base pairs, that implies nearly 200 mutations per base pair per human generation. It is in that sense that we meant above that the vast majority of possible alleles already exist at some level in the human population, and future evolution will be determined by a change in the frequencies of occurrence of various alleles. It is conceivable that if some presently rare alleles become much more frequent, new alleles might arise from multiple additional mutations built upon those presently rare alleles.
The concept of natural selection first introduced by Charles Darwin in his 1859 publication On the Origin of Species implies, in the language of modern genetics, that alleles that enhance fitness will naturally grow in frequency within a population. While fitness is measured ultimately by success in producing descendants, that success relies on many abilities: fertility; survival, in the face of possible predators, at least through the age of child-bearing (and for humans, as we will see later, possibly well beyond that age); resistance to disease; success in competing with other members of the species and with other species for possibly scarce resources needed for health and growth; success in attracting mates (to be dealt with in the next subsection); care and training of offspring; adaptability to a possibly changing environment. And all those abilities depend on a variety of phenotypic features of a species.
One can thus imagine an intricate fitness landscape, which can be envisioned in three dimensions as akin to the Herbert Bayer painting of Mountain Lakes pictured in Fig. II.1. The landscape features many peaks and many valleys, with peaks representing high fitness and valleys low fitness. The two dimensions of the painting surface would represent two specific phenotypic features, for example, eyesight and sense of hearing, or alternatively, brain capacity and means of mobility. Since fitness depends on many phenotypic features, the fitness landscape is truly multi-dimensional. But it’s easiest to visualize in three dimensions encompassing any pair of relevant phenotypic features.

Human evolution has led our species part way up one of these peaks in the multi-dimensional fitness landscape, while distant species have evolved up different peaks. Birds and bats acquiring wings led them onto a very different peak (in a decidedly different fitness landscape, since they must adapt to an environment that differs in many ways) than ours leading to bipedal motion with flexible arms and opposable thumbs. Octopuses climbed a very different peak concerning eyesight and nervous system distribution than did humans. Ancient branches in the phylogenetic tree of life corresponded to species separating to ascend different peaks from a common low starting point. More recent branches, such as that separating humans from chimpanzees, may represent only taking different paths up a common peak. The goal of natural selection is to lead a species toward the summit of their chosen local peak, even though it may not be the highest peak in the fitness landscape or correspond to the most highly developed specific phenotypic feature.
The fitness landscape peak humans are ascending is one corresponding to exceptionally large brain capacity, which allows us to outwit other species and to learn how to develop technology that aids our fitness (while also threatening our existence in some cases). That species choice also has influence on our reproduction and child-rearing. Human gestation periods are long while the fetal brain begins to develop in utero. But brain growth and development in utero is limited by the need for the baby to emerge through the vaginal canal. Further brain development takes many years, leading to an unusually long period of childhood care by parents and grandparents. And the long gestation and child care impose some limits on fertility of humans. But our large brains also allow us to exploit Earth’s resources so effectively that human population growth is also ultimately limited by the extent of those resources.
The majority of mutations that arise in new generations are either neutral or detrimental with respect to the fitness landscape. But natural selection works to favor those occasional mutations by which an allele leads a species further up their local fitness peak. The number of generations over which those helpful mutations may become frequent in the population depends on the strength of the fitness improvement they produce. But the structure of the fitness landscape and the nature of natural selection make radical evolutionary changes in phenotype highly unlikely. Radical changes correspond to different, possibly even higher, fitness peaks, so that to reach them would require some genetic changes that temporarily reduce fitness, in order to traverse a fitness valley. And such genetic changes are strongly disfavored by natural selection.
For example, humans might benefit from growing wings, but acquiring them would require changes to many genes, and each change would have to be lasting on its own through many generations – i.e., each would have to be either neutral or positively selected on its own, possibly for contributing at first to some improvement other than wings. Mailund puts it this way: “Selection gradually improves on what we already have; it does not make wild jumps from one form to another. If we have major changes in the future, they will evolve slowly and follow paths where each step improves on the previous state. We might see radical changes in our future, but they will take an extremely long time to come about…Each step evolution takes us cuts us off from places where other paths might have taken us.”
Often, genetic changes involve tradeoffs, increasing fitness in one way while decreasing it in another. As long as the increase is the stronger effect, the change will still be naturally selected. A good example is the sickle-cell allele of the hemoglobin gene, which gained frequency in populations exposed to mosquito groups that carried malaria. An individual who has a single copy of the sickle-cell allele has increased resistance to malaria and, hence, increased survival probability. However, individuals who receive the sickle-cell allele from both parents are then subjected to a different debilitating disease: sickle-cell anemia. Until now, malaria has been the greater threat in many parts of the world, so this allele has increased in frequency during human development. However, if modern medicine produces an effective cure or preventative for malaria, the sickle-cell allele will become negatively selected, and is likely to die out eventually.
The fitness landscape changes over time as changes in environment, disease exposure, diet, cultural pressures, evolution (or extinction) of competing or cooperative species, technology, etc. alter the conditions to which humans have to adapt. Thus, even as a species climbs its local fitness peak, the “ground” shifts beneath its feet, modifying natural selection pressures. Educated guesses about future human evolution are based on assessments of the changes in the fitness landscape from early human communities to the present densely populated globe. It is conceivable that unforeseen disasters, such as the aftermath of a giant asteroid collision with Earth, will cause “earthquakes” in the fitness landscape, leading to extinction (as it did for the dinosaurs some 66 million years ago) or radical evolutionary change in many species. But unforeseen (as opposed to foreseeable) disasters are beyond the scope of our discussion.
Sexual selection:
In order to produce descendants, one must first attract a mate. Thus, Darwin was led in his 1871 book The Descent of Man to add sexual selection as an important evolutionary pressure. Sexual selection has led quite spectacularly in other species to the development of male appearance and abilities that are especially attractive to females of the species, even though they offer the male no advantages – and may even prove disadvantageous – with respect to survival. The most obvious example is the elaborate tail feathers of the male peacock (see Fig. II.2), which would seem to send a look-at-me signal to potential predators. Birdsong and sometimes amusingly elaborate bird dancing rituals are sexually selected in other bird species. Such features seem to signal to females of the species that these are males with good genes to be passed along to offspring.

Scott Solomon points out that “Israeli ornithologist Amotz Zahavi noted that the male traits preferred by females are often exactly those that are detrimental to his survival. Zahavi’s idea, known as the handicap principle, suggests that such traits are honest indicators of good genes because males cannot fake these traits without the good genes to back them up…Females can therefore be assured that males with traits that essentially handicap them really are genuine.”
Male-male competition also factors into sexual selection. Traits that give males physical advantages in physical combat or competition with other males of the species over mate selection – the spurs on the legs of peacocks are an example — are also sexually selected.
Among humans, one obvious preference of most women is for taller men, and that may explain, via sexual selection, why there is a significant average height difference between human males and females. Solomon notes that in “a study of the U.S. Military Academy at West Point’s graduating class of 1950…taller men were shown to have fathered more children throughout the course of their lives than shorter men, in this case because the taller men remarried more frequently…suggesting that sexual selection did indeed favor taller men.” In terms of facial features that humans find attractive, studies suggest that greater facial symmetry is preferred. As with the brightness of a peacock’s feathers, greater than average height and facial symmetry among human males seem to signal better genes to potential female mates. Clearly, there has also been sexual selection in the body odors and pheromones characteristic of men and women.
And human males are not immune to Zahavi’s handicap principle of sexual selection. Young male aggressiveness and recklessness is believed to be attractive to many females, even while it threatens male survival. Those young males who survive risk-taking are often judged to have good genes by potential mates. However, the same traits are not attractive in fathers. Mailund notes: “When we are looking for a mate when we are young, we need to show off our genes. Once we have started a family, we need to protect our offspring. We can take risks to find a mate because we do not have much to lose. It is either finding a mate or not reproducing, so whatever it takes to look attractive is worth it. Once we have offspring, taking risks puts us at a selective disadvantage.”
Although attractiveness to potential mates often seems to be determined somewhat arbitrarily by chance or by culture, sexual selection pressure can be very strong. Mailund describes the phenomenon of Fisherian runaway: “Each generation has more of the attractiveness gene and more of the ‘desire’ gene, and very rapidly, the entire species will possess this particular trait and find the trait attractive. There will be an arms race for displaying more and more of the trait. We can see extremely rapid evolution this way, even if the trait is entirely arbitrary.” But at least among humans, perceptions of attractiveness can change with the age of females and males. Thus, as social and cultural pressures are leading women in developed countries to begin having children at later ages, traits that are sexually selected in males may change in the future.
Gene drift:
Natural and sexual selection both favor alleles that enhance evolutionary fitness. But sometimes, especially in small and isolated populations, random differences in birth rates among females in the early generations – differences that have nothing necessarily to do with genetic fitness – can lead to domination of allele frequencies in later generations. And women who do have genetically enhanced fertility and their mates may pass along to future generations, purely by chance, neutral or even harmful mutations and alleles that offer no particular adaptive advantage. Yet these alleles may still come to dominate the later generations.
Gene drift is the mechanism at work, for example, in the high rate of occurrence of some rare conditions among inbred populations, such as Ellis-van Creveld syndrome among Amish communities in America or total rod cell color blindness on the Pingelap atoll in Micronesia. It may also account for some of the branching seen in Fig. II.3 of early hominins into distinct species when they migrated from the African lowlands of human origin to form isolated groups elsewhere in Africa and in Eurasia.
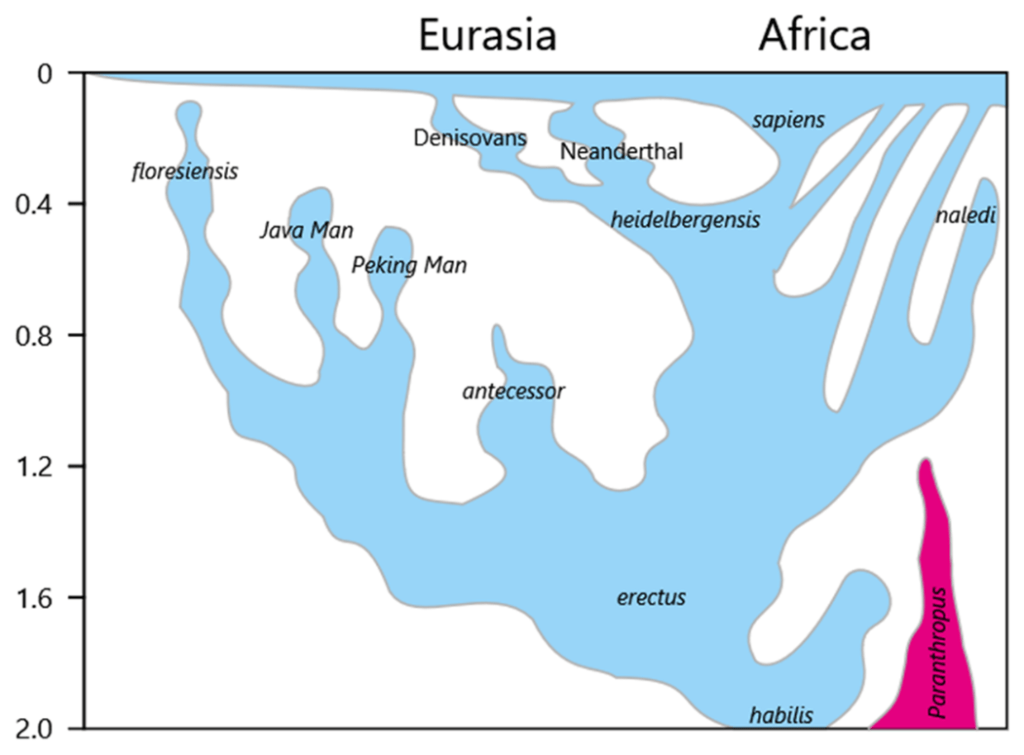
Gene flow:
Allele frequencies often differ significantly between separated populations, particularly when they adapt to different local environments. When migration and travel lead to mating between representatives of two such separated populations, evolution is advanced through gene flow from one population to the other. As migration and world travel have gotten easier in the post-industrial world, intermarriage between formerly distinct populations has led us toward a future in which humans around the globe will eventually share pretty much the same gene pool. To quote Scott Solomon, in that future “Any beneficial mutation that arises in one part of the world can quickly spread – but, so too, can new diseases.”
Historically, however, gene flow has played a significant role in human evolution to this point. It also involves gene flow between distinct species. In general, members of different species have difficulty mating with each other to produce viable descendants. For example, while male donkeys and female horses are often mated to produce mules, the mules are themselves sterile, so they can’t pass those combined genes onto future generations. But during the emergence from Africa homo sapiens did occasionally mate successfully with Neanderthals and Denisovans, and even with some still more ancient species within the genus homo, and many humans today still bear small imprints of the resulting gene flow in their DNA. Since the different hominin species tended to occupy different geographical locations, their gene flow into homo sapiens was also location-dependent, though subsequently spread out by further human migration, as suggested in Fig. II.4.
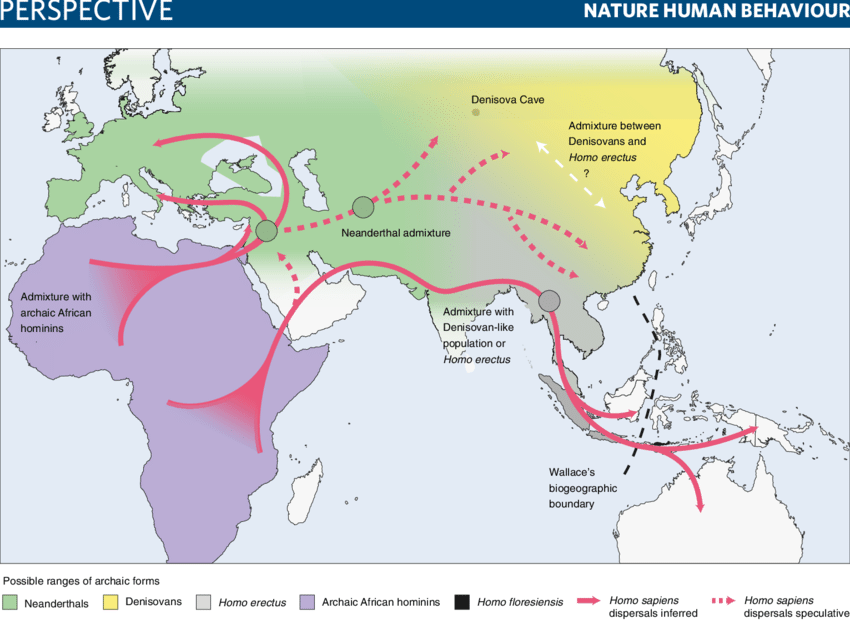
Human selection:
Modern humans have invented technologiesthat provide important aids to human adaptation and reproduction: disease cures and vaccines, prosthetic limbs, dietary supplements, pharmaceuticals to enhance brain function, in vitro fertilization, to name a few, leaving out the enormous array of advanced contemporary tools to aid human tasks. But these aids do not directly affect human evolution; we will discuss possible indirect effects of advanced human technology on future evolution in part II of this post. But the discovery in 2012 and rapid deployment since of CRISPR-Cas9 gene editing offers humans the technology now to change their own DNA. We have described CRISPR editing and some of its applications in a previous post.
The vast majority of CRISPR applications to humans to date involves gene editing in somatic cells, whose DNA is not passed on to offspring. Somatic cell edits are being used in clinical trials to attempt cures for genetic diseases, especially those like sickle-cell anemia that are reliably connected with a mutation within a single gene. They are also being used to enhance cancer immunotherapies. But even if the diseases treated are inherited, somatic cell edits only act on the treated individual and are not themselves heritable.
In order to have a direct impact on evolution, gene editing would have to be performed on germline cells – egg or sperm cells, or embryos – involved in reproduction. For now there is essentially a worldwide moratorium on germline editing in humans, especially in the wake of the 2018 announcement by Chinese entrepreneur He Jiankui that he had produced the world’s first “CRISPR babies.” He did this by removing the virus from sperm cells provided by two HIV-positive Chinese males, and then using those sperm for in vitro fertilization of egg cells provided by their HIV-negative wives. The fertilized egg cells were then CRISPR-edited to remove the gene that encodes a receptor protein for the HIV virus. This resulted in the birth of one set of twins who were allegedly then unsusceptible to later HIV infection or to passing HIV susceptibility along to future descendants. It turned out that the editing had been done sloppily, and in any case He Jiankui was subsequently sentenced to three years in prison and banned for life from working in reproductive science, for violating a Chinese regulation against germline editing in humans.
However, other Chinese researchers have continued to carry on CRISPR research involving germline editing on other animal embryos and on non-viable human embryos. Some of this research is aimed at removing susceptibility to heritable diseases, but some done in association with Chinese military institutions is aimed at exploring possible future enhancements of soldiers. One example that has received some press coverage is the editing of beagle embryos to remove myostatin-producing genes that normally limit muscle growth. A male and a female beagle resulting from these embryos, pictured in Fig. II.5, are said to have twice the muscle strength of normal beagles and to be considerably stronger and faster. Possible implications for human strength enhancement are clear.

We are thus on the threshold of a human selection era of evolution. The possibility of removing genetic diseases from entire lines of descendants – or even potentially from entire populations via the use of gene drives – is too enticing for scientists to ignore. An international commission formed from various national academies of science after the He Jiankui announcement produced a report in 2020 that supported “defining a responsible pathway for [future] clinical use of heritable human genome editing” when it is deemed to be both safe and medically necessary. The safety concern is important. At present, CRISPR editing still often ends up altering DNA elements beyond the targeted gene, and those unintended edits can have deleterious effects. Furthermore, we do not yet have a complete understanding of which genes contribute to which phenotype features, so even perfectly targeted editing may risk collateral damage to human functioning.
Further research to improve CRISPR technology and improved understanding of the impact of many genes will be needed before human germline editing becomes widespread. And human attitudes about its use may have a considerably longer path toward acceptance than the technological aspects. It seems inevitable, however, that human selection will be applied in the not-too-distant future, and most likely to enhance “desirable” phenotypic features as well as to eliminate some genetic diseases. Indeed, a preliminary form of human selection is already available to those who can afford it, in the form of preimplantation genetic diagnosis. In this technique, couples who produce multiple fertilized eggs in vitro can choose which one(s) to implant in the womb, discarding the rest, based on their preferences among genetic characteristics of the various embryos. Human selection and human gene editing have the potential to significantly alter the fitness landscape for natural and sexual selection, and must be taken into account in speculating on the future of homo sapiens.
III. humans have evolved
Hominins and chimpanzees formed separate evolutionary branches from common ancestors some 6-8 million years ago. We have documented the extensive similarities between human and chimpanzee genomes in a previous post. But it is worth mentioning here two of the most important differences. The gene SRGAP2, which encodes a protein involved in the formation of neurons and synapses during nervous system and brain development, underwent a series of duplications in hominins compared to chimpanzees. The multiple copies of this gene in humans lead to significantly denser neuronal structure, with more extensive interconnections, supporting greater cognitive abilities than in other mammals. In addition, the gene FOXP2 in Neanderthals, Denisovans and homo sapiens features mutations with respect to the corresponding gene among the apes and other mammals, and these mutations may have contributed to the development of speech and language.
The family tree of hominins as they developed after the split from apes, and culminating in the genus homo, is illustrated in Fig. III.1. In this section we will provide a number of examples of the further evolution of homo sapiens after they began to migrate out of their African birthplace some 50,000 to 100,000 years ago. Although the Neanderthals and Denisovans also migrated out of Africa, the fossil evidence suggests that only homo sapiens found ways to survive the last Ice Age, from which the Earth began to emerge about 20,000 years ago.
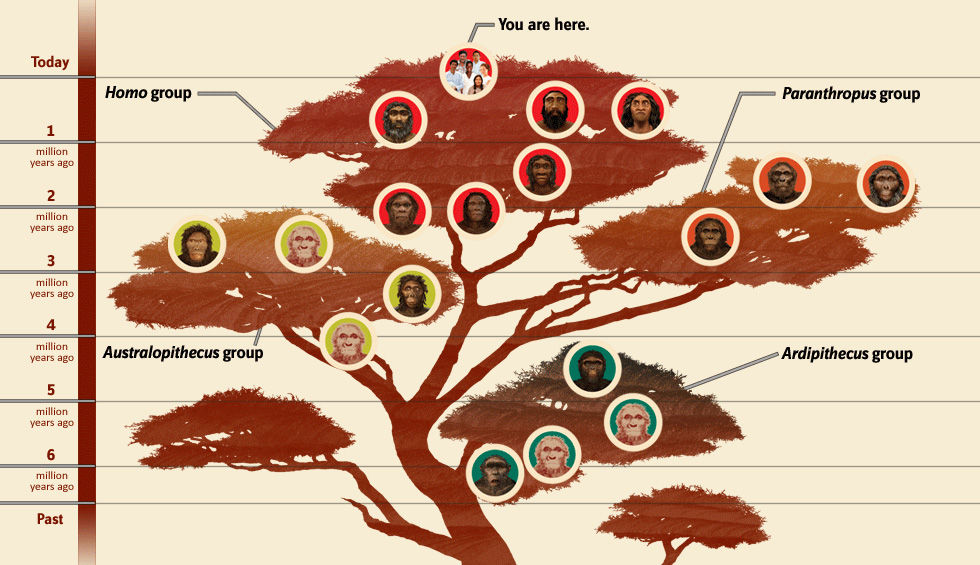
Adaptation to environment:
A number of evolutionary changes in humans since leaving the African lowlands resulted from natural selection pressure supporting adaptation to new environments and quite different climates. The most evident of these involves skin pigmentation (see Fig. III.2). When humans separated from the great apes and lost body fur, they became more susceptible to ultraviolet (so-called UVB) solar radiation, which was intense in the tropical lowlands of Africa where early hominins developed. UV radiation tends to destroy the B vitamin folate, which is required to make new copies of DNA, as needed especially for fetuses. Folate deficiency is linked to various birth defects. So reproductive efficiency among early humans required skin protection against UVB radiation.

This skin protection is provided by melanin, which is produced by skin cells called melanocytes. The melanocytes in individuals with darker skin make more of a darker version of melanin known as eumelanin. The eumelanin offers darker-skinned individuals even greater protection against UVB radiation, as was needed in the African tropics. However, the UVB light from the sun is needed to produce vitamin D in the liver and kidneys, and thereby the ability to absorb calcium and phosphate needed to build strong bones. As humans migrated to more temperate latitudes, where sunlight strikes the Earth more obliquely, the excess skin protection offered by eumelanin-fueled dark skin became a liability, because it inhibited vitamin D production.
So natural selection has favored genetic variants that led to lighter skin, as a better tradeoff between folate and vitamin D production. (Among modern humans, we are now seeing widespread vitamin D deficiencies attributed to both dietary changes and limiting exposure to sunlight to prevent skin cancers.) This is one example of the tradeoffs often involved in genetic adaptations to environment.
In particular, a single-nucleotide polymorphism (SNP) in a gene called SLC24A5 affects skin pigmentation and has been found to differ dramatically among different human populations. According to Scott Solomon, “Between 98 percent and 100 percent of Europeans have an adenine at the site [in question], whereas 93 percent to 100 percent of Africans, East Asians, and Native Americans have a guanine. Although other genes are now known to influence human skin color, SLC24A5 accounts for somewhere between one-quarter to one-third of the difference in skin color between Europeans and sub-Saharan Africans.” Allele frequencies have adjusted over many generations to differences in sunlight exposure in different locations. This is one of the dramatic allele differences between human populations that allow the company 23andMe to deduce a person’s geographical ancestry from their DNA samples.
We have previously mentioned the sickle-cell allele as another example where frequencies in different populations resulted from counter-balancing natural selection adaptation pressures. Evolution has made the sickle-cell allele, which distorts the shape of red blood cells, prevalent in parts of the world where malaria-bearing mosquitoes are abundant (see Fig. III.3), but rare elsewhere since the possibility of sickle-cell anemia has yielded negative natural selection pressure.

Additional examples of natural selection evolution are apparent in populations who live in high-altitude mountainous regions. But here isolated groups have developed distinct evolutionary mechanisms to deal with their common challenge of low oxygen pressure. Research has focused on three distinct and distant groups: Tibetans, natives who live in the high Andes, and the Amhara people of the Ethiopian highlands. Their different evolutionary adaptations and the genes involved in those adaptations are summarized in Fig. III.4. Oxygen is transported around the body by the hemoglobin in red blood cells. Thus, one way to compensate for reduced external oxygen pressure is to increase the concentration of hemoglobin in the blood.

Andean natives living at 13,000 feet, for example, have enlarged chest and lungs and high hemoglobin concentrations, leading them to have even more oxygen in their blood than do Peruvian natives at sea level. But this adaptation also has a downside: the high-altitude dwellers often suffer from chronic mountain sickness (CMS), which stems from having too many red blood cells. In contrast, Tibetans who live at high altitude have much less hemoglobin and oxygen in their blood than their Andean counterparts, but have developed to breathe faster. The Tibetans also have elevated levels of nitric oxide, which causes blood vessel dilation (as in an erection) and more capillaries than normal. Thus, they have elevated blood flow and more delivery sites for the reduced concentration of oxygen in their blood. The Amhara people of the Ethiopian highlands have hemoglobin concentration and arterial oxygen saturation not very different from Ethiopians living at sea level.
Genomic analysis of the Tibetans reveals some very interesting features, which are shared also with the Sherpas in Nepal. The affected genes are EPAS1 and EGLN1, which are both involved in sensing and reacting to the amount of oxygen that enters the body. The EPAS1 allele that helps Tibetans to adapt to low oxygen contains five distinct mutations not found in any other known human population. Surprisingly, the same five mutations to the EPAS1 gene have been found in the genome of the human-adjacent Denisovan population, in DNA analyzed from Denisovan skeletons. This similarity is unlikely to have occurred by random chance, and so it suggests that Tibetans have acquired that allele by gene flow from early humans in the Tibet region having occasionally mated with Denisovans, who themselves evolved after migrating from Africa to Asia (see Fig. II.4) a few hundred thousand years ago.
Adaptation to changing diet:
Those aspects of the human genome that affect digestion and storage of nutrients have evolved to date under natural selection dictated by the food supplies available to ancient humans. For example, the unreliability of access to animal protein and fats in early hunter-gatherer communities during the last Ice Age has led to a human need for and ability to store fat. There appear to be nearly a hundred genes that affect fat storage in humans. In today’s developed world, where fatty foods are abundant, that ancient natural selection pressure is leading to a high frequency of obesity, increasing susceptibility to conditions such as diabetes and heart disease. So we may well see the natural selection pressure for fat storage change in the future.
In the aftermath of the Ice Age, starting some 10,000 years ago, humans began to develop agriculture and the domestication of animals. That led to additional changes in diet, with natural selection impact on several specific genes. For example, the need in early agricultural communities to digest milk from domesticated cattle, sheep and goats has led to changes in the LCT gene that governs lactose tolerance. Lactose is the sugar contained in raw milk and, as indicated in Fig. III.5, the body’s ability to break down that sugar in the small intestine depends on the presence of an enzyme called lactase, whose production is controlled by the LCT gene. Mammals produce lactase when they are babies, in order to facilitate digestion of the mother’s milk. But in most mammals that production ceases after the babies are weaned.
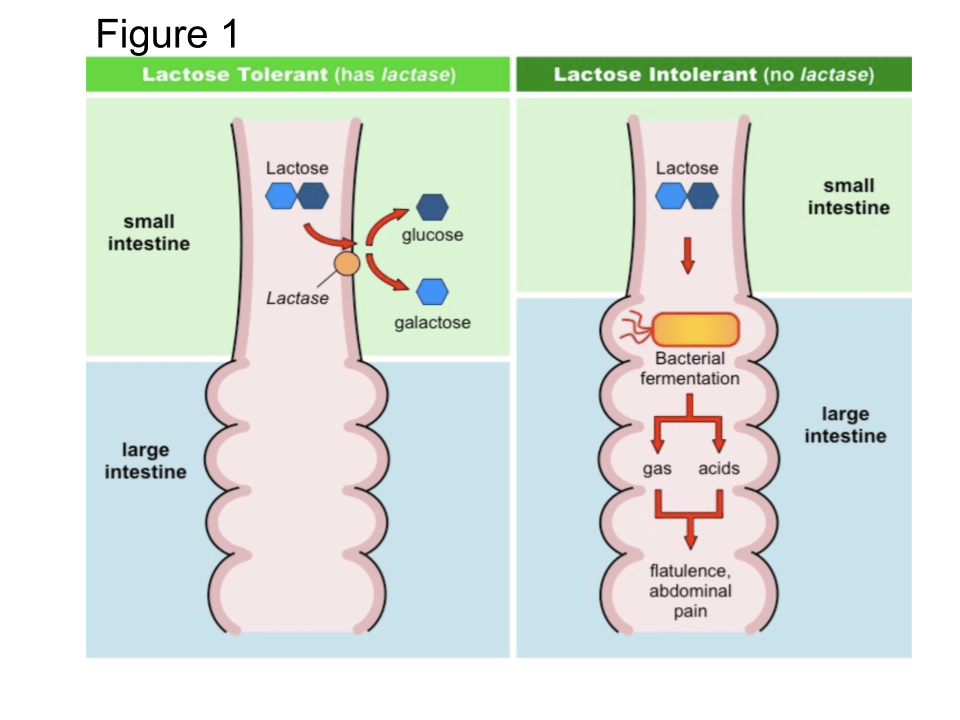
According to Scott Solomon: “People who can drink milk as adults simply never stop making lactase, thanks to a small evolutionary change in the genomes of their ancestors…A study using DNA from modern people as well as DNA extracted from ancient Eurasian skeletons found that the strongest signal for natural selection came from the LCT gene – and that the earliest appearance of the version of the gene that allows adults to digest milk was between 2450 and 2140 BC, suggesting that selection has been acting on this gene within the last four thousand years.” However, it has also been found that different mutations of the LCT gene were developed to enhance lactase production in northern European vs. North African and Middle Eastern pastoral populations, showing yet again that evolution often finds distinct genetic modification paths to favor the same adaptation.
Another digestive evolutionary adaptation concerns rice digestion, aided by an enzyme called salivary amylase that helps to break down the starch in rice already in the mouth. Modern humans have anywhere between 2 and 15 distinct copies of the gene that produces amylase, with more copies found systematically among populations like the Han Chinese and Japanese, where rice is a food staple requiring enhanced amylase production to avoid indigestion.
It should also be noted that the domestication of animals exposed early humans to new microbes that had previously been found only in the wild ancestors of those animals. And the increasing population density in early human civilizations made it much easier to spread the resulting infectious diseases to other humans. Scott Solomon again: “Indeed, the majority of modern infectious diseases – around 60 percent – are the result of a virus, bacteria, fungus or protozoan that switched from infecting an animal to infecting humans. We acquired anthrax from sheep and cattle; influenza from birds, pigs, and horses; rabies from dogs – and there are about two hundred other microbes that crossed species lines and began infecting people.” As we will see in part II of this post, one of the important avenues for future human evolution will likely come from natural selection pressure to improve resistance to such infectious diseases, of which the SARS-CoV-2 virus, and all its variants, are only the most recent example.
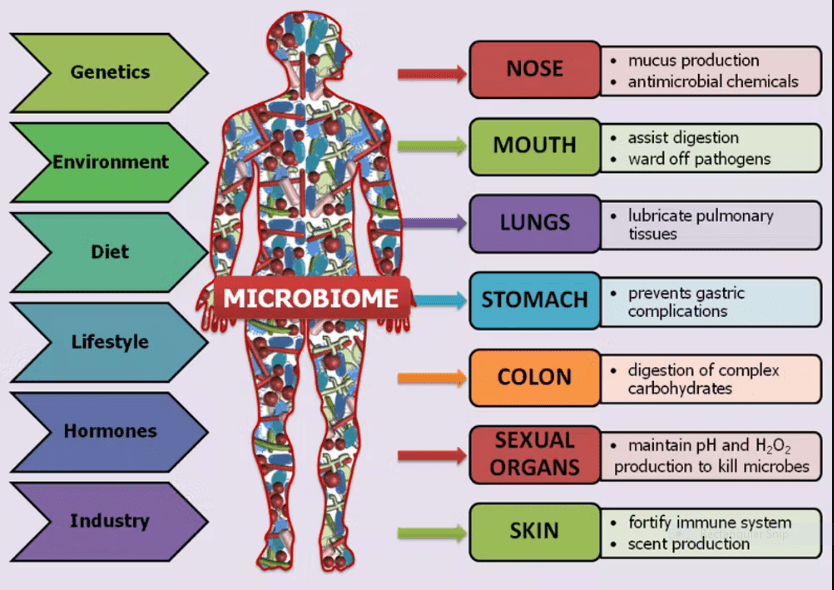
Not all microbes are harmful, however. Humans host a variety of microbes that aid various functions, including digestion, and that form a human microbiome (see Fig. III.6) that has come under genomic study only in the 21st century. Recent genome sequencing on bacteria found in the digestive tracts (mouth + intestine), skin and vagina of humans have provided evidence that humans and the human microbiome have evolved together (coevolution) over the last several million years. Scott Solomon: “…part of our evolutionary history involved outsourcing – some of the jobs that could be done in-house, by our own genes, are instead performed by the genes present in our microbial partners. This is an advantageous arrangement, in part because our microbiome contains one hundred times more microbial genes than does our own genome, meaning our partners can perform far more functions than we can on our own…our microbiome can evolve far faster than we can – about one hundred thousand bacterial generations for every one human generation. That means that in addition to swapping out one type of microbe for another, the microbes that remain with a particular host over many generations can quickly adapt as environments change. We need that versatility – conditions are changing more rapidly for people today than ever before in our history.”
However, the diversity and types of microbes found in human guts and teeth also depend strongly on diet and environment. Comparative microbiome studies of modern humans in western urban environments, in rural environments, and isolated jungle environments (e.g., the isolated hunter-gatherer Yanomami people discovered in the Amazon rainforest at the Venezuela-Brazil border regions), and of ancient human remains (particularly well-preserved teeth and fecal remains) seem to show systematically “that modern urbanites have lost a significant amount of microbial diversity, and some of the missing species might have been beneficial.”
Examination of human dental remains from Central Europe and the British Isles has revealed dramatic shifts in oral microbial communities following the development of agriculture and the Industrial Revolution. Some of these changes undoubtedly reflect changes in diet and sanitation, especially water quality. But some dwindling of the human microbiome also results from more recent changes in medicine, including both the rise of Caesarian sections that limit microbial transfer from mothers to newborns and the wide use of antibiotics. The collateral damage from antibiotics, in particular, has been highlighted in two books of the last decade: The Wild Life of Our Bodies: Predators, Parasites and Partners That Shape Who We Are Today by Rob Dunn and Missing Microbes: How the Overuse of Antibiotics is Fueling Our Modern Plagues by Martin Blaser. Microbiome losses have led to the multi-billion dollar global industry behind sales of probiotics. But they are also likely to lead to some future human evolution to replace lost functionality. We will discuss possible future evolutionary adaptations in part II of this post.
Figure III.7 summarizes some of the differences between ancient and modern human diets and associated health concerns and life expectancy changes.

Other historical human adaptations:
Natural selection favors fertility. It is reasonable, then, to wonder why evolution has led to human females living long past menopause, when they are no longer fertile. In this feature humans differ from the vast majority of other mammals, although there is some evidence indicating life beyond menopause among some non-human primates as well. Modern medicine clearly contributes to human life extensions, but there is also a likely evolutionary advantage related to the extensive child-rearing period needed during childhood human brain development.
Scott Solomon puts it this way: “Because they did not have to wait for their children to become self-sufficient before having another baby, [early] women whose mothers were helping provide food could have more children over the course of a lifetime.” Those grandmothers could also help in other ways to keep the children healthy until the children could produce offspring of their own. Thus, as child-bearing becomes more dangerous to the mother, as well as the baby, with increasing age, there is an evolutionary gain in allowing women to transition post-menopause to grandmother status. The natural selection pressure may have been particularly strong during the hunter-gatherer and early civilization stages, when average human life expectancy did not exceed 30 years. Those women who lived significantly longer than the average would have produced more children, and overseen the care of more children, who could then pass on those genes favoring longevity to future generations.
Sometimes allele frequencies undergo fairly rapid change when large fractions of a population are killed off by a disaster. Two such natural disasters struck Europe during the late Middle Ages, with devastating effect. The Great Famine of 1315-1317 is estimated to have caused the deaths of 10-25% of the human population in many cities and towns. This was followed in fairly short order by the Black Death caused by bubonic plague in Eurasia and North Africa from 1346-1353. During this short period, the plague is estimated to have killed 30-60% of the European population and about one-third of the population of the Middle East.
The survivors of these two great disasters, whether they were merely lucky or in some way possessed enhanced resistance through their genes, dominated future allele frequencies in the Western world. In particular, University of Colorado scientists who examined the remains of 36 later bubonic plague victims from a 16th-century mass grave in Germany have found that “innate immune markers increased in [allele] frequency in modern people from the town compared to plague victims.” Their study suggests “that these immunity genes may have been pre-selected in the population long ago but recently became selected through epidemic events.”
Examination of the types and causes of historical human evolutionary trends provides us with insights to now speculate on the future of human evolution in part II of this post.
