Steve Vigdor, June 11, 2020
This post belongs in the “brave new world” category, along with our earlier post on Gene Drives. In both cases, we are dealing with applications at the cutting edges of genomic research that raise controversial ethical and regulatory issues, but which are not yet sufficiently in the public eye to have attracted organized science denial. The issue I’ll deal with here resides at the interface of science, law enforcement, art, ethics, law and government. My own interest in the issue was, in fact, piqued by an art installation I saw in September 2019 at the Museum of Applied Arts in Vienna, Austria. But I’ll get to that art installation later in this post. My goal here, as it was for gene drives, is to discuss the underlying science, together with the promise, perils and controversy of its applications.
The central question at the heart of the technology to be described is how much of a person’s observable characteristics can be reconstructed from a small DNA sample or, in the relevant technical language (to be explained further below), how much of a person’s phenotype is determined from their genotype. How accurate is the information, and how much of it should be usable in forensic science in support of law enforcement and in courts? DNA evidence has, of course, been used in courts for decades, but we’re dealing here with a new type of DNA “evidence,” as will be explained in more detail below.
1. Genotype vs. Phenotype

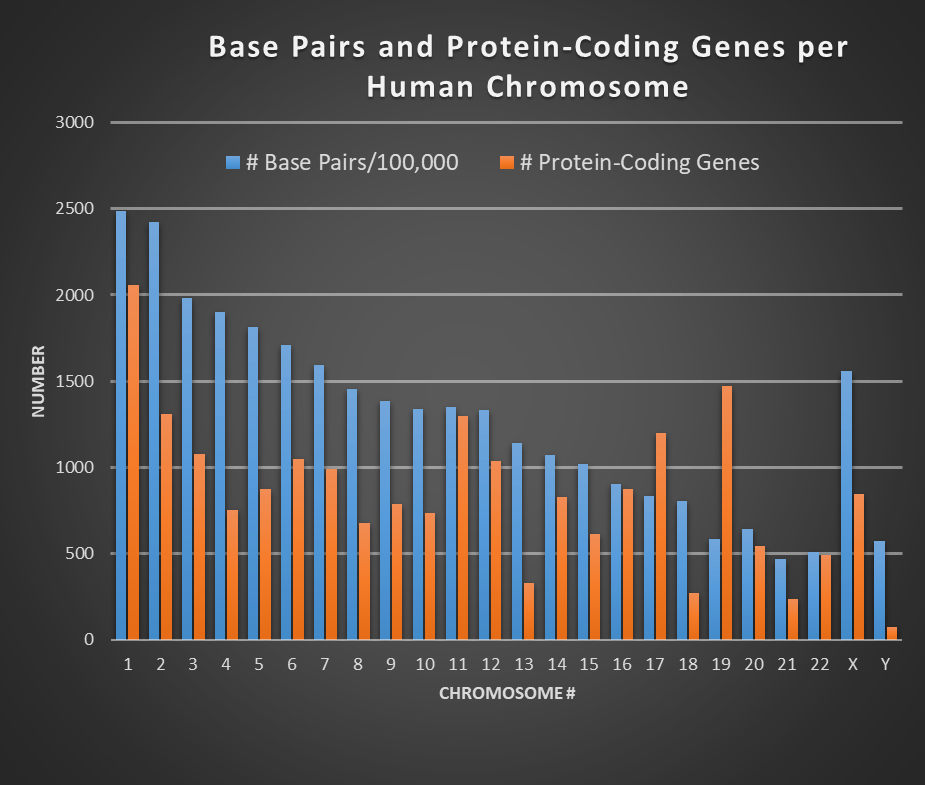
“Genotype” refers to the makeup and architecture of all the genes of an individual, as inherited from parents. Human DNA comprises 23 pairs of chromosomes, each containing many genes (see Fig. 2). In all, human DNA comprises roughly 30,000 genes or pseudogenes (non-functional DNA segments that resemble genes) among the 3 billion base pairs in the full human genome. The base pairs refer to the couplings of the four constituent nucleotides – adenine (A), thymine (T), guanine (G), and cytosine (C) – within the double helix structure of DNA. Among the genes, about 20,000 encode the assembly from amino acids of proteins essential for life functions. The coding genes account for only 1.5% of the total number of base pairs in the human genome. There are two copies of each gene, one inherited from each parent. Many genes have different variations, called alleles. An individual will often inherit different alleles of a specific gene from the two parents. Some alleles are dominant and some recessive.
“Phenotype” refers to the ensemble of all outwardly observable characteristics of an individual, including form and structural features, physical appearance, development and behavior. The genotype plays an important role in determining features of the phenotype, but is not the sole contributor. Environment can also play an important role and can, for example, affect the expression of genes. Even identical twins, who share exactly the same genotype, can have subtle differences in appearance, have different fingerprints, and so forth. These can arise from inherited epigenetic differences, which do not involve differences in the sequences of nucleotides within the DNA, but rather in developmental or chemical processes that alter the expression of genes. Furthermore, of course, physical appearance can be altered by hair style, hair dye, facial hair, cosmetic surgery, tanning, etc. The scientific question at the heart of forensic DNA phenotyping concerns the accuracy with which phenotypic features can be determined from a mapping of the genotype from a very small DNA sample, for example, left behind inadvertently at a crime scene.
Only about 0.1% of human DNA differs among individual humans. The rest determines our common characteristics as members of the same species. However, 0.1% of 6 billion total (including both parental chromosomes) base pairs still leaves about 6 million base pairs that can determine individual differences. There is generally not a one-to-one mapping of genes into phenotype characteristics. Rather, individual features of appearance – eye color, hair color, skin pigmentation, facial shape, hairline, etc. – are typically influenced by multiple genes, and by the pairing of alleles for those genes. Ongoing research is trying to understand those influences by analyzing DNA samples from very large numbers of individuals, and searching (most recently with analyses applying machine-learning computational techniques) for increased frequency of specific alleles among those who share certain observable traits.
The analysis of the specific DNA sections that differ among individuals is generally done by using polymerase chain reaction (PCR) techniques. The PCR process was developed by Kary Mullis in 1983 and led to his share of the 1993 Nobel Prize in Chemistry. PCR mimics the natural process of DNA replication, but limits it to specific DNA sequences of interest, because those contain the differences among individuals. The application of PCR allows the DNA section of interest, drawn from a tiny sample, to be rapidly reproduced millions or billions of times to create an amplified sample suitable for study. The analysis can be done quite rapidly using modern (so-called Next Generation) sequencing techniques that allow many small DNA fragments to be read out in parallel.
2. Uses of dna in forensic science

The analysis of individually differentiated DNA samples for forensic purposes is done in two quite different ways. The traditional method is known as DNA profiling or DNA fingerprinting. This approach normally concentrates not on the genes themselves, but on sections in between coding genes in up to 20 specific DNA regions, looking at so-called short tandem repeats (STRs). These are short (typically, 3 to 5 base pairs long) DNA sequences that are repeated different numbers of times in different individuals. For example, labeling the four nucleotides that form DNA by T, A, C and G, a repeated sequence might be four adjacent sites with TATT. That sequence might be repeated three times at one location in one individual and five times in another. A profile is constructed from the number of repeats of each such STR in each of 20 different DNA regions. The chance that two people who are not related share identical STR profiles is theoretically about 10–18, i.e., one part per billion billion. In practice, the identification uncertainty is considerably worse than this, and is typically dominated by laboratory error and DNA sample contamination. Still, DNA profiling can be used reliably to see if DNA scrounged from a crime scene matches a suspect’s DNA. This is the type of DNA evidence that is often used in court cases to demonstrate that a suspect was present at the crime scene.
The new method – the subject of this post – is called DNA phenotyping. This is an attempt to predict at least some features of an unknown suspect’s phenotype from a small sample of DNA collected at a crime scene. This approach focuses not on STRs, but rather on single-nucleotide differences that occur at specific DNA locations, either within protein-coding genes or in DNA sections that affect gene expression. These differences are called single-nucleotide polymorphisms (SNPs). The greatest interest is in SNPs within genes that effect changes in the protein sequence coded by the gene, and thus correspond to different alleles of the gene, as in Fig. 4, where a CG pair in one allele is replaced by an AT pair in the second allele. More than 335 million SNPs have been found across humans from multiple populations. Some SNPs are much more common in certain geographic groups than in others, and that is the basis for the now popular DNA-sequencing ancestry determinations.

Certain SNPs in specific genes are associated with an individual’s susceptibility to some particular diseases, such as sickle-cell anemia or cystic fibrosis. Others may affect how individuals respond to specific pathogens, chemicals, drugs or vaccines. Such associations are critical to personalized medicine, where treatments are tailored to individuals based on their genetic predisposition to respond to treatments or disease risks.
Connections of specific SNPs to specific aspects of phenotype are established statistically through genome-wide association studies (GWAS), in which DNA sequencing is done for large cohorts of individuals who share a certain trait, for comparison to control groups who do not share that trait. The GWAS are “fishing expeditions” in which the entire genome (not just selected sections) is analyzed in search of SNPs that occur with significantly higher frequency in the cohort as compared to its control group. For example, a 2005 GWA compared genomes for 96 patients with age-related macular degeneration of eyesight to those for a control group comprising 50 healthy individuals. The study found two specific SNPs with significantly altered allele frequency between the two groups.
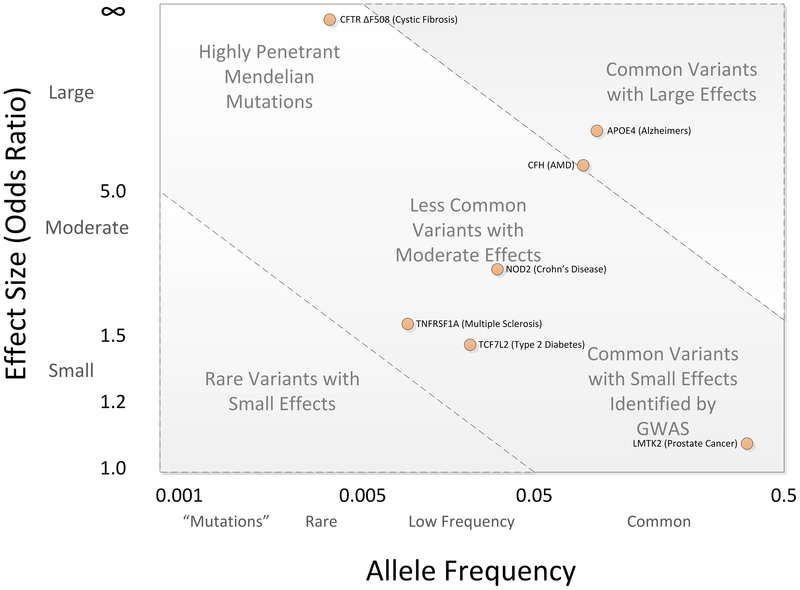
By now, several thousand disease-related GWAS have been performed, and currently they often include cohorts containing a million or more individuals, to better establish the statistical significance of allele frequency differences. Figure 5 shows for some diseases the degree of correlation between particular alleles and susceptibility to the disease (the odds ratio of getting the disease for individuals with, relative to those without, the allele of interest) on the y-axis vs. the frequency of the correlated allele in the general public, on the x-axis. So, for example, cystic fibrosis is quite rare among humans, but is caused by mutations to a specific gene. Prostate cancer is very common, and there is a weak correlation in allele frequency between prostate cancer sufferers and control groups.
3. biogeographic phenotyping
The popular geographic ancestry determinations that are now routinely offered by companies such as 23andMe or AncestryDNA are based on a collection of SNPs that differ in allele frequency between different populations. There are roughly 15 million SNP sites within the human genome, among which some can be selected as ancestry-informative markers (AIMs). High-frequency SNPs tend to get passed down through generations in a specific location and become relatively common in a given population. Sets of AIMs are selected for providing the highest accuracy as a marker for a specific geographic ancestry. For example, North and South Han Chinese ancestry can be distinguished unambiguously using a set of 140 AIMs, while nearly all sub-Saharan Africans have one particular allele that occurs very infrequently in populations outside this region. In general, the accuracy of biogeographic phenotyping is quite good at the continent level (Europe, sub-Saharan Africa, East Asia, South Asia, Oceania and Americas), but less accurate with regard to sub-continental origins, as ancestral migration within continents lessens the predictive power of the AIMs. However, five combinations, or clusters, of AIMs have been identified that allow one some degree of discrimination of ancestors’ geographic origins within Europe, as shown in Fig. 6.

In biogeographic phenotyping, the Y chromosome in males and mitochondrial DNA in females are of particular importance, because they and their contained AIMs are passed down essentially unchanged over many generations within the male and female lineages, respectively. Their analysis thus allows some separation between the geographical ancestry inherited from a person’s father and mother. On the other hand, among the other 22 pairs of chromosomes, only one member of each pair comes from each parent, meaning that only half of the parents’ full complement of AIMs are typically passed along to each offspring in each generation. Consequently, phenotyping is generally of limited accuracy in inferring mixing of ancestors from different geographical regions, if that mixing occurred many generations before.
4. reconstructing facial appearance
In forensic DNA phenotyping, the goal is to make a predictive model of the appearance and biogeographic ancestry of an unknown crime suspect from sequencing of a DNA sample left at a crime scene. The focus in applications to date has been on reconstructing the facial appearance from DNA. As we will see below, this is an inexact science so far, but it has already been used in multiple criminal investigations in both North America and Europe.
By now, predictive models have been generated for eye color (which has the advantage that the bulk – though not all – of phenotypic variation between brown and blue eyes can be traced to SNPs in a single gene) and for hair color among European populations. 41 different SNPs have been identified as linked to hair, eye and skin color in humans. 17 SNPs are currently used to determine most likely skin pigmentation. 15 loci in different genes have been recently identified as linked to facial shape features, including the shape of the nose. The distinctions so far are relatively crude in most models. For example, eye pigmentation is judged to be most likely blue or brown or intermediate; hair color, blond, brown, red or black; hair shade (light or dark); skin pigmentation, very pale, pale, intermediate, dark, and dark to black. But research is ongoing and models are likely to improve rapidly.
So, how are facial reconstructions actually done? Once a small DNA sample is available, PCR is used to select the subsections of interest and to amplify the sample. The amplified sample can be sent to a commercial DNA genotyping laboratory (e.g., 23andMe or Illumina) to get back the nucleotide sequence map of the DNA subsections of interest. This map can then be analyzed with software that compares the sequence to stored SNP templates for various facial features. That software can then provide a model “mug shot” based on the “average” appearance of individuals who share the same gender and biogeographical ancestry as the subject, with modifications to that average appearance determined from the most probable associations of SNPs with eye color, hair color, skin pigmentation, and other facial features.
Such facial reconstruction modeling is now offered commercially by several companies. Most prominent to date has been Parabon NanoLabs, a Virginia-based company that offers Snapshot DNA Phenotyping Service, based on proprietary software that creates composite face imaging sketches based on DNA samples. This is the modern genomic take on traditional suspect sketches that were based on eyewitness descriptions. The U.S. Department of Defense has provided approximately $2M in development financing for Snapshot. Other companies that offer analogous services are Illumina, Inc. and the Swiss company Identitas AG.
DNA phenotyping has so far been used mainly as a way of trying to locate suspects in cold cases. When sufficient DNA samples, clearly identified with the crime itself, are recoverable from a crime scene, they are first used for STR profiling, in order to compare with DNA profiles that may be stored in an investigative database or a publicly accessible database like GEDMatch, and that may be obtained from an eventually identified suspect. (In general, customers for services such as those provided by 23andMe are given an option to allow their de-identified genotype information to be used for research purposes. However, the 23andMe “fine print” includes the following disclaimer: “…we do not share customer data with any public databases, or with entities that may increase the risk of law enforcement access. In certain circumstances, however, 23andMe may be required by law to comply with a valid court order, subpoena, or search warrant for genetic or personal information.”) If there is sufficient sample remaining after the STR analysis, then investigators can try to do phenotyping to inform probabilistic estimates of the biogeographical origins and the facial features of a not-yet identified suspect.
The construction of phenotyped “mug shots” has by now been used successfully (i.e., in a way leading to an arrest confirmed by subsequent STR profile comparisons with the identified suspect) to identify suspects in a handful of cold cases in North America and Europe. As of April 2019, the suite of Parabon’s genetic genealogy tools – including, but not limited to, Snapshot – has helped lead to 49 suspect identifications, about 1000 renewed investigations, and to at least 17 arrests. Among these are previously unsolved murders in 1986 (Tacoma, Washington), 1988 (Fort Wayne, Indiana), 1996 (Idaho Falls, Idaho), 2009 (Lake Charles, Louisiana), and 2011 (Columbia, South Carolina).
The Idaho Falls case is worthy of note because a previous suspect had spent 20 years in jail for the crime, despite the fact that his DNA did not match that collected at the crime scene. He was finally released in 2017 after the DNA phenotyping was completed. And a new suspect identified with the aid of phenotyping was arrested, and confessed to the crime, in 2019. There were also two rape and murder cases in the 1990s in the Netherlands solved with the aid of DNA phenotyping. Snapshot composites have been released publicly in several other unsolved cases, including ones of sexual predation, in the hopes that they might lead to identification of suspects.
There is also at least one example of an earlier crime, eventually dated to 1963, in which DNA phenotyping for both facial features and geographic origins has been used in conjunction with facial reconstruction from a recovered skull to identify a previously unidentified victim.
A great deal of controversy has been generated in Germany by the so-called Bavarian Police Task Act, which passed the region’s legislature in May 2018. This act permits police use of DNA phenotyping not only in solving past crimes, but also to identify suspects who might represent an “imminent danger” to others. Specifically it permits the “molecular genetic examination of found trace material…for the purpose of determining the DNA identification pattern, sex, eye, hair and skin colour, biological age and biogeographical ancestry of the person causing the trace […] if averting the danger would otherwise be hopeless or significantly more difficult.” Clearly, there is the possibility of discriminatory abuse in such preemptive actions, where no crime has yet been committed, and the concerns go to the heart of the controversy surrounding forensic DNA phenotyping. We will return later in this post to a detailed discussion of concerns about phenotyping.
5. predictive value of dna phenotyping
What is known about the accuracy of phenotyped “mug shots”? The lion’s share of the facial reconstructions used to date in forensic studies has been carried out with the proprietary software of Parabon NanoLabs, for which there are not yet peer-reviewed analyses of accuracy. Indeed, Parabon has been criticized for this lack of openness: Moses Schanfield, Professor of Forensic Sciences at George Washington University has criticized the lack of peer review, noting that at the time he wrote there was no publicly available performance record for Parabon’s Snapshot product. The American Civil Liberties Union in 2016 recommended using DNA phenotyping only “…where the link between genes and external characteristics is based on well-proven, peer-reviewed, widely accepted science, such as is apparently now the case with hair and eye color.”
The Parabon website makes the qualitative claim that “using deep data mining and advanced machine learning algorithms in a specialized bioinformatics pipeline…the Snapshot Forensic DNA Phenotyping System…accurately predicts genetic ancestry, eye color, hair color, skin color, freckling, and face shape in individuals from any ethnic background, even individuals with mixed ancestry.” However, the reports they provide to clients have included a number of disclaimers, for example, that their “prediction models do not represent the full range of human genetic diversity,” that environmental factors “can affect appearance in ways that are inherently unpredictable,” and that “discretion should be used when attempting to include or exclude individuals in an investigation by comparison of appearance with Snapshot predictions.“
The Parabon website does now feature a poster with results shown in Fig. 7, reporting a blind study of accuracy of Snapshot reconstructions of 24 volunteers for a University of North Texas (UNT) project done in collaboration with Parabon. The subjects were recruited by UNT based on their phenotypic and ancestral diversity. 25 anonymous DNA samples were sent to Parabon, including one two-person mixture. Each sample was genotyped using an Illumina chip sorting through 851,274 SNPs, and the results were run through the Snapshot algorithm to reconstruct skin color, eye color, hair color, prevalence of freckles, width, height and depth of facial features, regional ancestry and a composite facial reconstruction for each blind sample.
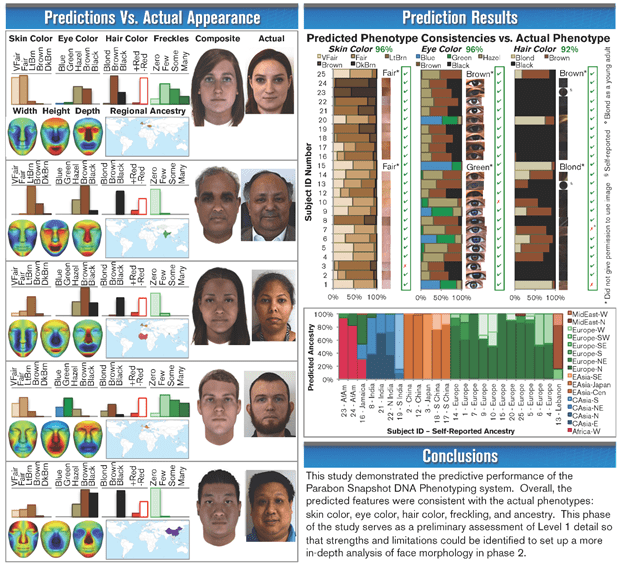
Note that the Snapshot algorithm provides more detail on both facial features and biogeographical origins than is currently agreed to be accurately accessible from the SNPs. Furthermore, the composite reconstructions are based on an average appearance for individuals from the same gender and ancestry groups, using the detailed SNPs from the genotype to then evaluate the subject’s differences from that average. The phenotypic reconstructions for Fig. 7 were done assuming a default age (25 years) and body mass index (BMI=22) for each subject. Age and BMI values for each subject were then sent to Parabon after the phenotypic reconstructions, and the facial composites were then age-progressed by a forensic artist. Figure 7 compares those age-progressed composite facial reconstructions with photographs for 5 of the subjects, as well as Parabon predictions with self-reported phenotypic characteristics and ancestry for each of the 25 DNA samples.
Based on the limited sample in Fig. 7, the UNT team reports accuracy of the Parabon reconstructions to be 96% on both skin color and eye color (one error on each among the 25 samples) and 92% on hair color. Although there is no comparable quantitative accuracy stated for geographical ancestry, the results look quite accurate in comparison with self-reported ancestry of the subjects. It should be kept in mind, however – and we will return to emphasize this point in the following subsection – that the algorithm produces probabilities that each subject falls into each of the allowed categories for eye color, hair color, skin color, and ancestry region. The composite reconstructions represent what the Parabon algorithm decides is the most probable look, but one could also reconstruct less probable, but still viable, composites based on phenotypic estimates of lower probability.
Much more statistically meaningful analyses of forensic DNA phenotyping predictive value have been carried out for various publicly available phenotype reconstruction algorithms. Among these public algorithms, the most extensive one to date, based on data drawn from multiple European populations, is the HIrisPlex S system developed in a collaboration between a group at Erasmus MC University Medical Center in Rotterdam, the Netherlands and the Walsh laboratory at Indiana University Purdue University Indianapolis, USA. This algorithm predicts eye, hair and skin color from a total of 41 single-nucleotide polymorphisms among the 22 pairs of non-sex-determining human chromosomes. They provide the table below of current prediction performance metrics for various phenotypic traits, deduced from the standard deviations obtained in 1000 comparisons of most probable reconstructions from DNA phenotyping with actual visible characteristics of individual subjects in their database.

The table above presents five different metrics to characterize predictive value for the phenotyping algorithm for each of the visible facial characteristics considered. The two most relevant metrics to measure the forensic usefulness of DNA phenotyping are the positive (PPV) and negative (NPV) predictive values. The PPV represents the fraction of subjects whose genotype contains positive markers for a given visible characteristic who actually exhibit that characteristic. In other words, it is the ratio of true positive predictions for a characteristic to all positive predictions. In the same way, NPV is the ratio of true negative predictions to all negative predictions (genotype indicates absence of the visible characteristic in question). The PPV and NPV can be quite different from each other. For example, genetic predictions of an eye color intermediate between blue and brown (e.g., hazel) are of very limited usefulness, but genetic indications that the eye color is not intermediate have a 91% chance of being correct.
The table shows that the predictive value of current DNA phenotyping models varies widely among specific visible characteristics. At best, DNA phenotyping currently has about 90% probability of getting blue eye color and dark-to-black skin color correct. In contrast, the probability of getting brown hair color or pale skin color correct is only about 2/3. It is critical to keep these probabilities in mind when evaluating DNA phenotypic composite sketches, which represent some sort of average “best guess” at a subject’s appearance.
The accuracy of specific predicted features does not quite address the central question for forensic use of these composites, which is their usefulness in identifying potential suspects, if those suspects are not at all known to start with. The New York Times, in reporting on early uses of DNA phenotyping in cold case investigations, tested this question in an interesting, but non-scientific, way. They did not use the Parabon Snapshot software, per se, but rather used publicly available software for facial reconstruction from phenotyping developed by Mark Shriver, a professor of anthropology and genetics at Penn State, and Peter Claes of KU Leuven in Belgium. Parabon has said that their proprietary software is based in part on Shriver’s work.
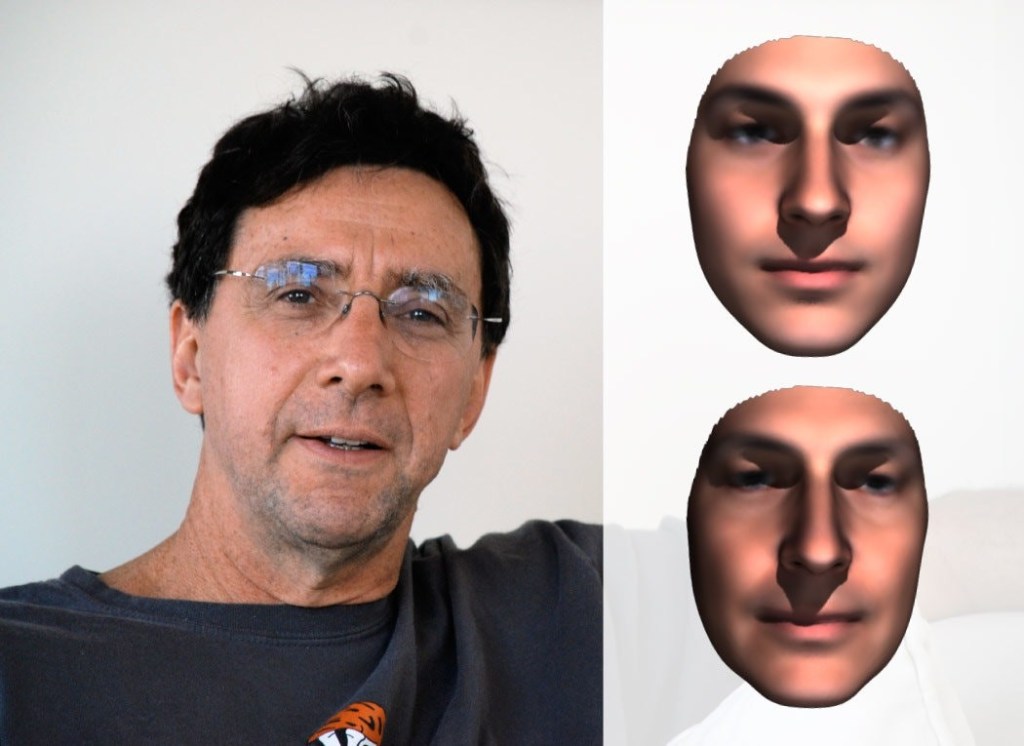
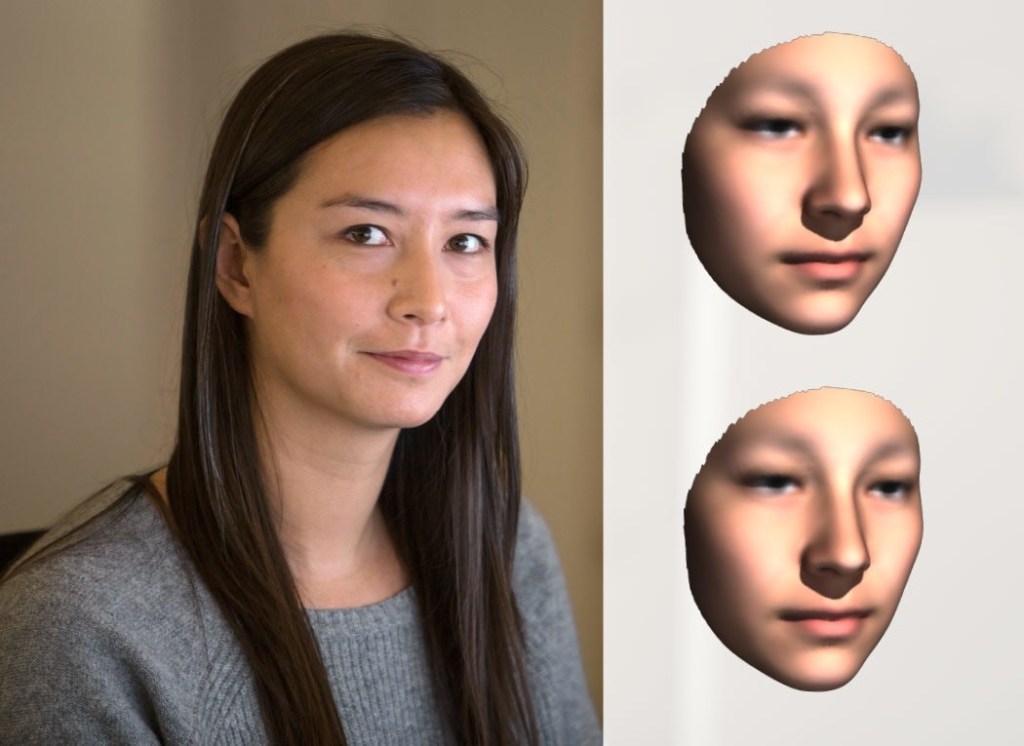
The Times’ science desk submitted to Shriver the genotypes of two employees who volunteered their 23andMe results. Shriver then did phenotyping to produce facial reconstructions of the two, without any knowledge of their names, ages, heights or weights. The Times then asked colleagues of these volunteers if they could identify their colleagues from the facial reconstructions, while being told that the person in each case could be younger or older, or lighter or heavier, than Shriver’s phenotypic images suggested. None of the 50 or so Times colleagues of both subjects who ventured guesses correctly identified the male subject from the DNA phenotyping facial reconstruction, while 10 correctly identified the female subject.
Photographs of both subjects are shown side-by-side with Shriver’s facial reconstructions in Figs. 8 and 9. In each case, the lower right composite was produced after the fact, when Shriver corrected for the then available information on age, height and weight of the subjects. When shown side-by-side with photographs, the facial reconstructions look pretty good, if slightly generic, but it is challenging to identify a subject, even among a confined pool of possibilities, from the reconstruction alone.


6. probably chelsea
The ambiguity in DNA phenotyping reconstructions was stressed in the art installation that triggered my interest in this subject. The work, entitled Probably Chelsea, was created by artist Heather Dewey-Hagborg as a result of a commission she was given by Paper Magazine to produce a composite of Chelsea Manning to accompany a 2014 mail interview the magazine published with Manning. Manning is the transgender U.S. Army Intelligence analyst-turned-whistleblower who went to prison for sharing classified information about the prevalence and scope of civilian deaths and torture in the Afghan and Iraq wars. At the time of the interview, the imprisoned Manning could not be visited or photographed.
Dewey-Hagborg, who had previously created images of strangers from found DNA, received some mouth swabs and hair clippings from Manning. She used PCR herself to extract and amplify the DNA sections of greatest interest, had them sequenced by a commercial company, and then used her own custom facial reconstruction software to do the phenotyping. Her software was adapted from publicly available forensic facial recognition software used in Basel, Switzerland. She then used 3D printing to produce full-size full-color “masks” of the reconstructed face.
Manning presents a unique challenge to DNA phenotyping, which normally considers gender a characteristic easily fixed from a person’s genome (is there or is there not a Y-chromosome?), since she had undergone hormone therapy to transition from male to female. For the portrait Dewey-Hagborg produced for the Paper Magazine interview and displayed as the art installation Radical Love, she then generated two versions of Manning’s face, one “female” and one androgynous, as shown in Fig. 10. As Dewey-Hagborg herself explains: “Placing these two portraits side by side I made apparent the reductionism of pinning someone’s gender to simplistic readings of genetic sex—a routine practice in DNA forensics. … Even biological or genetic sex, commonly considered to be simple and straightforward turns out to be amazingly complex. Genetic pathways related to secondary sexual characteristics and hormone production are scattered around the genome on various chromosomes and many remain unknown. These phenotypes vary on a spectrum, are mutable and show the limits of efforts to use DNA to predict gender.” She references the 2013 book by Sarah Richardson, titled Sex Itself: The Search for Male and Female in the Human Genome.

But gender is far from the only characteristic subject to genotypic ambiguity, and Dewey-Hagborg subsequently decided to embrace this ambiguity fully. For Probably Chelsea, she took DNA phenotyping probabilities seriously, and produced 30 distinct versions of Manning’s possible appearance by choosing various phenotypic features from genotypic possibilities of lower than maximum probability. For example, again quoting Dewey-Hagborg:
“Chelsea’s mitochondrial DNA is special and at the same time it is totally ordinary. Other DNA variations tell similarly complex stories. For example, the GG variant of her rs12913832 polymorphism, which is often considered synonymous with blue eyes in Northern Europeans, is also found in Hispanic, African American, and South Asian populations, with varying phenotypes. So the same exact data can be read in different ways. This variant might predict she is most likely to have blue eyes and be of European ancestry, but there is still a good chance she could have brown eyes and she might not have much or any European ancestry at all.
Each genomic variation is a piece of data, a new clue and another possible story. As more data is put together some things become more probable, and some less, but there is never certainty and there are always alternate possible narratives. Probably Chelsea portrays these alternate narratives and represents a sampling of the many stories Chelsea’s DNA can tell.”
The resulting art installation Probably Chelsea was first exhibited after President Obama commuted Manning’s sentence and she was released from prison. She attended that first exhibition opening. The work has by now been exhibited in a number of different galleries and museums around the world. My own photo of the installation at the Vienna Museum of Applied Arts is pictured in Fig. 11, overlaid with an excerpt from the poster describing the piece. Probably Chelsea provides a haunting visualization of the possible, fairly wide, variations on DNA phenotyping, with neither gender, nor skin pigmentation, nor eye color, nor facial structure being definitively fixed by the genotype.

7. ethical concerns about forensic use of dna phenotyping
Given the considerably less than perfect predictive value of DNA phenotyping, and the ambiguity captured in Probably Chelsea, what applications of such phenotyping should be allowed and what applications should be forbidden? This question is the subject of ongoing debate in both Europe and the U.S.
Clearly, the accuracy is not great enough to be used in court proceedings, unless an indictment aided by DNA phenotyping has produced a subject whose DNA STR profile matches that of the crime scene sample. As an investigative aid, phenotypic composites should perhaps be compared to the more traditional composite subject sketches produced from eyewitness descriptions. In the U.S., the Innocence Project has revealed that 71% of more than 360 erroneous verdicts, identified after conviction by STR profile analysis, had been reached, at least in part, on the basis of false eyewitness descriptions and identifications. At least with DNA phenotyping it should be possible to estimate the probability of misleading features in constructed composite sketches.
The debate over police use of forensic DNA phenotyping has been most pointed to date in Germany, due to the adoption of the 2018 Bavarian Police Task Act. The major areas of concern are: the potential misuse of DNA phenotyping for racial profiling or discrimination against certain minority groups; the invasion of privacy by release of protected (e.g.,medical) susceptibility information accidentally included within the analyzed DNA sections; conflict with laws protecting the confidentiality of certain information; and exaggerated expectations of “scientific accuracy” on the part of the public, of juries, and of users of the DNA phenotypic information.
For example, phenotyping conclusions about biogeographical ancestry are often mistakenly taken as implying “ethnicity” or “race,” neither of which are determinable from DNA analyses. But the misinterpretation can lead to biased composite sketches based on misleading “average” characteristics for individuals of given gender and ancestry. This possibility is exacerbated by the fact that police investigations often seek to narrow down the search for subjects by focusing on relatively rare visible deviations from such average characteristics, while scientifically, the rarer the characteristic, the lower the probability of accurate determination, because the reference database is limited.
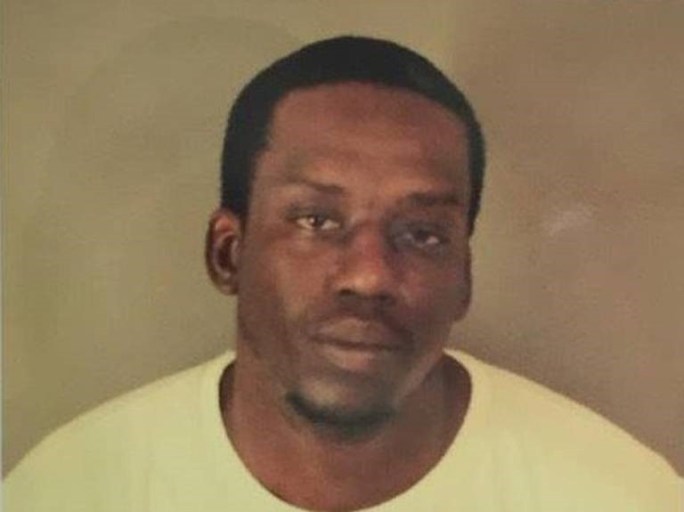
The concerns about racial profiling were brought to a head by the first public release in 2015 of a Parabon DNA Snapshot of the suspect in a cold (2011) double homicide case in Columbia, South Carolina. The Snapshot, shown in Fig. 12, basically depicts an “average” male of West African descent. It could have been used as a premise for questioning enormous numbers of African-American males. The composite sketch is compared side-by-side in Fig. 12 with a photograph of the suspect subsequently charged with the homicides in 2017 after a positive DNA profiling match to samples collected from the crime scene.
The release of the Parabon “mug shot” in Fig. 12 stimulated an article by Heather Dewey-Hagborg in The New Inquiry. There she points out that biogeographical ancestry determined from DNA phenotyping assesses “an individual’s percentages on roughly four ‘ancestral’ types: African, European, Native American, and East Asian; a division which recapitulates the centuries-old racial [and eugenics] categories of Caucasian, Mongoloid, and Negroid… Though race doesn’t exist at the genetic level, scientists can still create it by correlating large genetic datasets into classic racialist categories. This desire to find correlations pointing to race while simultaneously denying its [genetic] existence is what sociologist Troy Duster calls the ‘molecular reinscription of race.’”
Dewey-Hagborg goes on:
“While a recent justice department ruling banned racial profiling for federal law enforcement, it remains a common policy among local police, border patrol, and airline security screeners. It is not difficult to imagine how the vague racial images and descriptions FDP produces (ie. ‘dark skinned 92 percent african 8 percent european male’) could be used by police as a ‘scientifically legitimate’ excuse to justify racial dragnets. Furthermore, given the general low level of mathematical and scientific literacy in the population at large, it seems unlikely education of the end-users (police) will be sufficient to enable adequate interpretation of this complex information, its probabilistic nature, and the inherent uncertainties involved… These technologies black-box their stereotypes, hiding them in the algorithms. If you or I, or the police, or a corporate service enter DNA data into software and get racial percentages as output, the act of stereotyping is displaced. It isn’t us, it’s the system. The process doesn’t appear biased because the bias is coded in the program.”
In addition to concerns about racial profiling, there are issues related to confidentiality. For example, European Union data protection regulations place genetic data under special protection because they can reveal highly personal information, e.g., about a person’s mental or disease dispositions. Bavarian legislators attempted to prohibit the use of such personal characteristics, as opposed to information about “outwardly recognizable characteristics.” But, as pointed out by two German legal experts: “Unlike a computer code, the characteristic data in the DNA is not stored separately. Rather, there are many interrelationships, some of which have not yet been sufficiently researched, especially if the so-called “biogeographical ancestry” is to be identified. The more meaningful this finding should be, the more snips in the DNA must be analyzed. It can never be ruled out that these may also be relevant for the most sensitive [personal information].”
The same paper makes a distinction between the possible use of DNA phenotyping as one tool to narrow down a list of suspects in a long-running investigation, as opposed to its intended use in the Bavarian Police Task Act for “averting imminent danger.” “Dangers are usually urgent. Combating them requires rapid and valid knowledge. Forensic science and DNA phenotyping in particular cannot provide this… If a trace of DNA found on an explosive, for example, is found in a database, this makes it possible to identify the so-called ‘dangerous person’ or a person in his or her environment. This does not require phenotyping. The only possible statement [from DNA phenotyping] would be that the ‘endangered’ or persons from his or her environment may have certain external characteristics or that their ancestors could come from a more or less identifiable region of origin. Under perfect conditions this may be different in the case of longer-term criminal investigations, for example if the number of suspects can be reduced using such probability information.”
8. the legal situation
Few clear legal restrictions have been enacted yet to regulate uses of forensic DNA phenotyping. In a recent review, Schneider, Prainsack and Kayser have summarized the legal situation in Europe: “As of December 2019, forensic DNA phenotyping is explicitly regulated and permitted by law in two EU member states (the Netherlands and Slovakia), and practiced in compliance with existing laws in seven more (the United Kingdom, Poland, the Czech Republic, Sweden, Hungary, Austria, and Spain)… In Switzerland, it is forbidden under current law, but the legalization of forensic DNA phenotyping is currently being considered. In Germany, in November 2019, the Bundestag (Parliament) and Bundesrat (Federal Council) approved a change in the law to permit forensic DNA phenotyping (with the exception of the DNA-based inference of biogeographic ancestry…).” In the U.S. FDP is being applied with increasing frequency and little government regulation to date.
What regulations could be considered? In 2016, the American Civil Liberties Union’s senior policy analyst Jay Stanley suggested the following guidelines for the use of FDP:
“In some circumstances it may make sense to use genetic phenotyping in an investigation, but:
- Only where the link between genes and external characteristics is based on well-proven, peer-reviewed, widely accepted science, such as is apparently now the case with hair and eye color. [My caveat: see the predictive value estimates in the HIrisPlex S table above, even for hair and eye color.]
- Only where such evidence is used in conjunction with other, more specific evidence. It may be helpful to narrow down suspects, but it is not specific enough, and should never be used, to create suspects where none are otherwise known.
- It should not be used to pressure people to ‘voluntarily’ provide DNA samples or to submit to any other intrusive investigatory techniques. We don’t want police using these genetic analyses to run around conducting dragnets for every person in a neighborhood with black hair, hazel eyes, and freckles, for example.
- It should not be considered to constitute probable cause. Even in the case of a DNA sample that reveals that a suspect has albinism or some other particular genetic condition—perhaps a rare one—that fact should not be used to pour through hospital records, for example.”
The two German legal experts have added suggestions about transparency and certification of FDP results:
“In order to avoid open or even covert discrimination, a transparent scientific procedure is needed that makes it possible to question the results in a qualified manner. The transparency must relate to the reference databases and the evaluation methods used. A certification procedure for the evaluation processes should be defined. Added to this are regulatory mechanisms that minimize or eliminate the potential for discrimination associated with this investigation approach. In order to avoid misinterpretations, it must be ensured that the investigating officials receive comprehensive training and advice; the results of the investigation must show that they come from a genetic analysis and indicate calculated probability values. Public searches on this basis must be ruled out.”
We can expect more frequent use of FDP and more public discussion of its promise and its limitations over the coming years. I will conclude with another quote from Heather Dewey-Hagborg. While she is an artist, she shares concerns with many scientists involved in genomic research, but she manages to articulate those concerns more eloquently and pointedly:
“If we are indeed entering a future of genetic surveillance, it is the complexities, limits, biases, and weaknesses of these new technologies we need to excavate. To do so, we need a multifaceted and transdisciplinary approach blending art, science, theory, and hands-on experimentation. The media will talk about how it all works, but to fully understand, to appropriately educate others, to devise suitable policies, and to form strategies of resistance, we need to know how it breaks.”
Indeed, there is an opportunity, given the rapid pace of genomic research, to understand how and how often the assumptions underlying DNA phenotyping break down, before its misapplication leads to too many erroneous convictions. There are well-documented cases where the accuracy of previous forensic science techniques – e.g., hair comparisons, physical characteristics associated with arson fires, comparative bullet lead analysis, lie-detector tests – has been oversold in court and led to wrongful convictions. According to the Innocence Project, such misapplied forensics have contributed to 45% of wrongful U.S. convictions subsequently demonstrated by DNA profile mismatches. DNA phenotyping can be a helpful technique in crime-solving, but only if applied with a clear-eyed understanding of its limitations.
references:
https://en.wikipedia.org/wiki/DNA_phenotyping
H. Machado and R. Granja, Emerging DNA Technologies and Stigmatization, in Forensic Genetics in the Governance of Crime (Palgrave Pivot, Singapore, 2020), pp. 85-104, https://link.springer.com/chapter/10.1007/978-981-15-2429-5_7#enumeration
P.M. Schneider, B. Prainsack and M. Kayser, The Use of Forensic DNA Phenotyping in Predicting Appearance and Biogeographic Ancestry, Deutsches Arzteblatt International 116, 873 (2019), https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6976916/
B. Kaelin, DNA Phenotyping: Revealing the Faces of Killers, https://www.forensicsciencedegree.org/dna-phenotyping-revealing-the-faces-of-killers/
M. May, Next Generation Forensics: Changing the role DNA plays in the justice system, http://sitn.hms.harvard.edu/flash/2018/next-generation-forensics-changing-role-dna-plays-justice-system/
https://en.wikipedia.org/wiki/Genotype
https://en.wikipedia.org/wiki/Allele
https://en.wikipedia.org/wiki/Phenotype
https://en.wikipedia.org/wiki/Epigenetics
https://en.wikipedia.org/wiki/Polymerase_chain_reaction
https://en.wikipedia.org/wiki/Kary_Mullis
https://en.wikipedia.org/wiki/DNA_sequencing#Next-generation_methods
https://en.wikipedia.org/wiki/DNA_profiling
https://en.wikipedia.org/wiki/Single-nucleotide_polymorphism
http://www.personalizedmedicinecoalition.org/Userfiles/PMC-Corporate/file/pmc_age_of_pmc_factsheet.pdf
https://en.wikipedia.org/wiki/Genome-wide_association_study
R.J. Klein, et al., Complement Factor H Polymorphism in Age-Related Macular Degeneration, Science 308, 385 (2005), https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1512523/
https://en.wikipedia.org/wiki/Macular_degeneration
W.S. Bush and J.H. Moore, Genome-Wide Association Studies, PLoS Computational Biology 8, e1002822 (2012), https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1002822
https://en.wikipedia.org/wiki/Ancestry-informative_marker
M. Bauchet, et al., Measuring European Population Stratification with Microarray Genotype Data, American Journal of Human Genetics 80, 948 (2007), https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1852743/
https://en.wikipedia.org/wiki/Mitochondrial_DNA
https://hirisplex.erasmusmc.nl/
P. Claes, et al., Genome-Wide Mapping of Global-to-Local Genetic Effects on Human Facial Shape, Nature Genetics 50, 414 (2018), https://www.nature.com/articles/s41588-018-0057-4
https://www.illumina.com/services.html
https://snapshot.parabon-nanolabs.com/
https://www.gedmatch.com/login1.php
A. Mak, Genetic Genealogy’s Less Reliable Cousin, Slate, July 25, 2019, https://slate.com/technology/2019/07/parabon-nanolabs-genetic-genealogy-phenotyping.html
R. Boone, Idaho Man Who Didn’t Match Murder DNA Freed After 20 Years, The Spokesman-Review, March 22, 2017, https://www.spokesman.com/stories/2017/mar/22/idaho-man-who-didnt-match-murder-dna-freed-after-2/
E. Shapiro, Man Murdered in 1960s Identified Through Genetic Genealogy, ABC News, June 19, 2019, https://abcnews.go.com/US/man-murdered-1960s-identified-genetic-genealogy-years-people/story?id=63805623
C. Momsen and T. Weichert, From DNA Tracing to DNA Phenotyping – Open Legal Issues and Risks in the New Bavarian Police Task Act and Beyond, https://verfassungsblog.de/from-dna-tracing-to-dna-phenotyping-open-legal-issues-and-risks-in-the-new-bavarian-police-task-act-pag-and-beyond/
M. Schreiber, Police Turn to New DNA-Powered Technology in Hopes of Finding Killer, Washington Post, Feb. 23, 2015, https://www.washingtonpost.com/national/health-science/new-technology
J. Stanley, Forensic DNA Phenotyping, https://www.aclu.org/blog/privacy-technology/medical-and-genetic-privacy/forensic-dna-phenotyping
R. Wiley, et al., Blind Testing and Evaluation of a Comprehensive DNA Phenotyping System, https://docs.parabon.com/pub/Parabon_Snapshot_Scientific_Poster-ISHI_2016.pdf
https://walshlab.sitehost.iu.edu/
H. Murphy, I’ve Just Seen a (DNA-Generated) Face, New York Times, Feb. 23, 2015, https://www.nytimes.com/2015/02/24/science/dna-generated-faces.html?auth=login-email&login=email
https://anth.la.psu.edu/people/mds17
https://www.kuleuven.be/wieiswie/en/person/00041773
https://deweyhagborg.com/projects/stranger-visions
https://deweyhagborg.com/projects/radical-love
H. Dewey-Hagborg, Probably Chelsea, https://deweyhagborg.com/content/6-projects/10-probably-chelsea/probably-chelsea-essay.pdf
S.S. Richardson, Sex Itself: The Search for Male and Female in the Human Genome (University of Chicago Press, 2013), https://www.amazon.com/Sex-Itself-Search-Female-Genome/dp/022608468X
G. Samuel and B. Prainsack, Forensic DNA Phenotyping in Europe: Views “on the Ground” From Those Who Have a Professional Stake in the Technology, New Genetics and Society, 2018, https://www.researchgate.net/publication/329314232_Forensic_DNA_phenotyping_in_Europe_views_on_the_ground_from_those_who_have_a_professional_stake_in_the_technology
https://www.innocenceproject.org/eyewitness-identification-reform/
C. Cookson, DNA: The Next Frontier in Forensics, Financial Times, Jan. 30, 2015, https://www.ft.com/content/012b2b9c-a742-11e4-8a71-00144feab7de#axzz3Tt5OSqf0
T. Kulmala, California Man Held in 2011 Slayings of Columbia Mother and Daughter, The State, March 30, 2017, https://www.thestate.com/news/local/crime/article141735144.html
H. Dewey-Hagborg, Sci-Fi Crime Drama with a Strong Black Lead, The New Inquiry, https://thenewinquiry.com/sci-fi-crime-drama-with-a-strong-black-lead/
T. Duster, A Post-Genomic Surprise. The Molecular Reinscription of Race in Science, Law and Medicine, The British Journal of Sociology 66, 1 (2015), https://onlinelibrary.wiley.com/doi/epdf/10.1111/1468-4446.12118


