October 25, 2021
I. introduction
From time to time the harnessing of a new technology changes the way we live and the way we wage war profoundly. In the middle of the 20th century, the exploitation of fission and fusion processes revealed in research on reactions of nuclei led to the development of nuclear energy and nuclear weapons. In the late 20th century, the miniaturization of electronic circuits and the development of the internet paved the path to today’s world of smart phones, personal computers, social media, cybercrime and cyberweapons. The military and espionage applications of both of these technologies have led to arms races in which various countries try to gain offensive or defensive advantages over their adversaries, resulting in “mutually assured destruction” standoffs.
The 21st century has already seen the development of a new technology whose impacts over the long term may be even more profound than those of nuclear and cyber technologies. CRISPR gene editing, discovered in 2012, offers the promise of curing genetic diseases, contributing to the treatment of cancer and altering the evolution of species that serve as vectors for severe human diseases. At the same time it creates fraught opportunities for enhancing the capabilities of a select few humans and for development of new biological weapons that can alter the nature of warfare. Military and defense departments in the governments of major countries are well aware of those possibilities and we are currently in the infancy of a CRISPR arms race, which in some ways mirrors an evolutionary arms race between bacteria and viruses that has provided the insights to drive basic CRISPR research.
The development of CRISPR editing has a number of aspects in common with the development of nuclear technology and one very important aspect in common with the digital revolution launched in the late 20th century. As was the case for nuclear fission, the technology grew out of basic, curiosity-driven laboratory research, which led in very short order to both investigations of military applications and Nobel Prizes for the discoveries. The nuclear fission process, in which a heavy nucleus can be induced by neutron bombardment to split into two or more lighter fragments, releasing a great deal of energy, was discovered in 1938 by Lise Meitner and Otto Hahn, with Meitner’s nephew Otto Frisch helping with the interpretation of the experimental results. The potential implications of this process for both energy production and weaponry were immediately clear to many. The Manhattan Project to develop an atomic bomb based on nuclear fission of uranium was formally launched in the U.S. in 1939, only a year after the discovery. Hahn (but not Meitner!) won the Nobel Prize in Chemistry for the discovery in 1944.
CRISPR complexes were first discovered in basic research mapping the genome of various bacteria and other single-celled organisms, which had evolved over billions of years primitive immune systems to fight off viral infections. The detailed structure of the CRISPR complex and the mechanism by which it could destroy the DNA of invading viruses were first demonstrated by Jennifer Doudna of UC Berkeley and Emmanuelle Charpentier (then at the University of Umeå in Sweden, now at the Max Planck Institut in Berlin) in 2012. They also indicated how such CRISPR complexes could be engineered to modify essentially any DNA sequence of interest. For their work launching the CRISPR editing revolution Doudna and Charpentier (see Fig. 1) were jointly awarded the 2020 Nobel Prize in Chemistry.

Immediately after the discovery by Doudna and Charpentier, several groups worldwide advanced to demonstrate by the beginning of 2013 the use of a CRISPR complex involving the associated DNA-cleaving enzyme labeled Cas9 specifically to edit human cells. Gene editing had been possible before the CRISPR demonstrations, but the discovery dramatically improved the ease, efficiency and accessibility of doing so. As was the case for fission, the potential applications for both curing disease and creating advanced bioweapons or genetically enhanced soldiers were immediately clear after the discoveries. And again, only a short time passed before the military got involved in funding research into such applications. In the U.S. the Safe Genes program was launched by the Defense Advanced Research Projects Agency (DARPA) in 2016. And China’s Academy of Military Medical Sciences was not far behind.
However, the material and the technology needed to construct a nuclear weapon is very challenging and still, after eight decades, available only to a handful of state actors. The important feature that CRISPR editing shares with the digital revolution is that it is relatively cheap and widely available. Do-it-yourself (DIY) CRISPR-Cas9 kits, complete with samples and instructions for genetically altering bacteria, are available online for prices in the range of $100-300 US. You don’t have to be a molecular biologist to learn to use the technology relatively quickly. Community biology laboratory space is available for citizen scientists and hobbyists to carry out CRISPR experiments. The-Odin site which sells CRISPR kits was started by biohacker Josiah Zayner, whose stated goal is “to make the genetic engineering revolution as open and crowdsourced as the early digital revolution was.” Zayner has said: “We now all have this ability to program life. If millions of people took it up, that would immediately change medicine and agriculture, contributing so much to the world. By demonstrating how easy CRISPR is, I want to inspire people to that.”
Of course, the ubiquity of small computing systems in homes, businesses and government offices, combined with the widespread understanding of how to hack computer code and exploit its vulnerabilities, has led not only to enhanced contributions to worldwide knowledge and efficiency, but also to the viral spread of deadly misinformation and to the cyber warfare we have discussed elsewhere on this site. And that warfare is carried out now not only by nation-states, but also by “democratized” DIY hackers. Might we not expect a similar development with widespread CRISPR biohacking? Do we have to worry now not only about the proliferation of human and bioweapon enhancements via CRISPR carried out by the most developed nation-states, but also by DIY’ers not constrained by the Biological Weapons Convention? Cyber warfare can disrupt operations and threaten critical infrastructure, but genetically modified new biological weapons can be instruments of genocide.
This post has two parts. In the first part, in order to give a flavor for the pace of technical development, we discuss the basic science of CRISPR editing and review the history and current status of its development, ongoing research and clinical applications. In Part II we explore the worldwide military interest in CRISPR applications and discuss the ethical quandaries raised by the development of the means for humans to alter their own genome. Some of those quandaries were already suggested in Aldous Huxley’s novel Brave New World decades before even the structure of DNA was revealed.
II. a brief history of crispr development
The basic research leading to the development and advancement of CRISPR gene editing, together with compelling biographies of some of the key players, has been brilliantly described for a lay audience in Walter Isaacson’s recent book The Code Breaker: Jennifer Doudna, Gene Editing, and the Future of the Human Race. Another excellent source is Jennifer Doudna’s own book (written with Samuel Sternberg) A Crack in Creation. Here we summarize only a few of the most important highlights of that history.
CRISPR was first discovered in basic research toward the end of the 20th century to map the genome of various bacteria and other single-celled organisms known as archaea. A recurring pattern found in a number of species involved clusters of repeated DNA sequences separated by short normal-looking DNA segments referred to as spacers. One of the researchers working on the archaea genome, Francisco Mojica, coined the acronym CRISPR to describe these “clustered regularly interspaced short palindromic repeats.” The spacers were subsequently found to match DNA sequences found in certain viruses. And bacteria that contained CRISPR sequences featuring spacers that matched genomic segments in a particular virus seemed to be immune to infection by that virus.
This immunity was confirmed by two researchers, Rodolphe Barrangou and Phillipe Horvath, working for yogurt culture developers interested in defending their critical bacteria from viral infections. Barrangou and Horvath furthermore demonstrated that the acquisition of appropriate spacers by bacteria and their effectiveness in resisting viral infection relied critically on a CRISPR-associated (Cas) enzyme protein labeled Cas9; when Cas9 was removed from the bacterial DNA, the resistance to viral infection was lost.
These discoveries led to a race to understand the detailed structure of the CRISPR complex, the mechanism by which it operated to disable invading viruses, and whether it could be used for some purposes in human cells. The winners of this race, Charpentier and Doudna, formed a transatlantic collaboration in 2011. It had already been shown that Cas9 was essential as the molecular scissors to cleave the genetic material of the invading virus. Charpentier had demonstrated that a small snippet of RNA – known as CRISPR RNA, or crRNA for short – matching part of one strand of an invading virus’ DNA, was essential to guide the Cas9 to cut at the relevant location within the viral genome.
Charpentier and Doudna together demonstrated that one more critical piece of the complex was another RNA section – labeled as trans-activating CRISPR RNA, or tracrRNA – that served both to assemble the crRNA in the first place and then as a kind of handle supporting the entire complex as it latched onto the invading virus, so that the crRNA could then direct the Cas9 protein to cut the viral genome at the desired location.
Doudna and Charpentier published their results in June 2012. But they not only reported the structure of the CRISPR-Cas9 complex – they also stated the vast possibilities opened by their discovery. As Doudna has said: “Once we figured out the components of the CRISPR-Cas9 assembly, we realized that we could program it on our own. In other words, we could add a different [engineered] crRNA and get it to cut any different DNA sequence we chose.” Doudna and her team furthermore demonstrated that they could engineer a single RNA molecule – called single-guide RNA, or sgRNA — to combine the functions of crRNA and tracrRNA, making the CRISPR-Cas9 complex “an even more versatile, easy-to-use, and reprogrammable tool for gene editing.” They concluded their paper by pointing out that this CRISPR-Cas9 complex with engineered sgRNA, could be a more versatile and easily applicable alternative to existing gene-editing methods in humans.
Doudna and Charpentier shared the 2020 Nobel Prize in Chemistry “for the development of a method for genome-editing,” with the press release noting that “This technology has had a revolutionary impact on the life sciences, is contributing to new cancer therapies and may make the dream of curing inherited diseases come true.” But Doudna and Charpentier had performed their experiments on bacteria, not on human cells, where getting the CRISPR complex to penetrate into the nucleus of targeted human cells in order to modify their DNA is a much trickier business. So their paper launched a worldwide race to demonstrate that ability to work in human cells. This was accomplished by the early months of 2013, primarily in three labs: Doudna’s at Berkeley, Feng Zhang’s at the Broad Institute of MIT and Harvard, and George Church’s at Harvard. Figure 2 shows individual photos of these four key players, Church, Doudna, Zhang and Charpentier. Church had earlier supervised the Ph.D. research of both Doudna and Zhang.

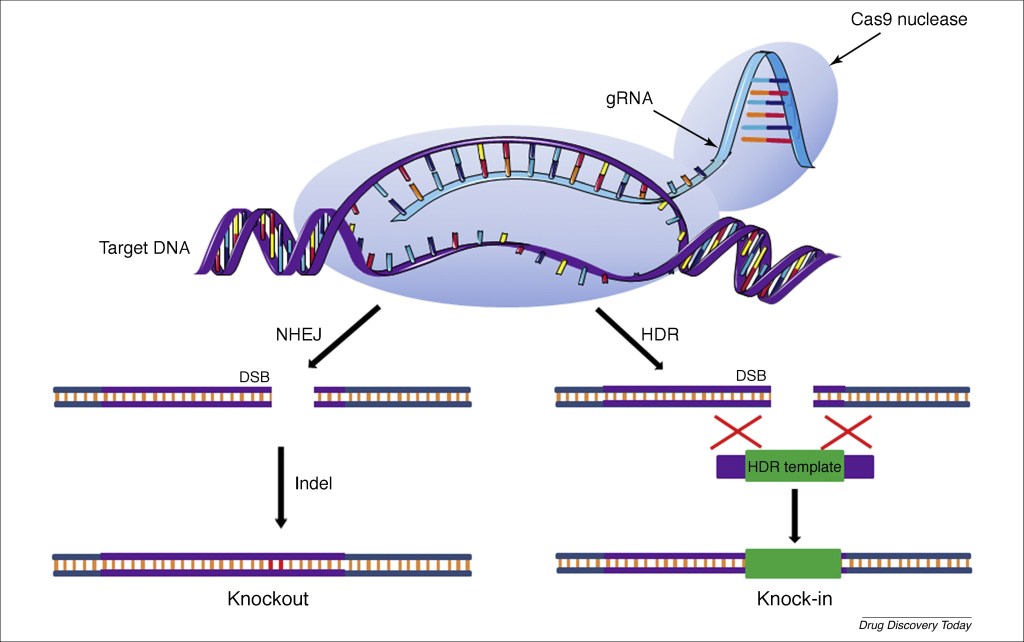
Figure 3 shows the entire CRISPR complex and how it operates. The crRNA section of the guide RNA (gRNA) attaches to one complementary strand of the targeted section of DNA, and the Cas9 enzyme then cleaves both strands (forming a double strand break, DSB) adjacent to DNA marker sequences known as PAMs (protospacer adjacent modif). After the cleavage, the break is repaired by one of two natural DNA repair mechanisms: either “non-homology end joining” (NHEJ) for gene knockout, or homology-directed repair (HDR) to copy a nearby available template section for precise gene replacement (or knock-in).
Concurrent with the demonstrations of CRISPR-Cas9 editing in human cells, the leaders of this research saw the immediate need to form one or more biotech companies that could oversee the development of therapeutic applications. At first all of the founders discussed joining forces in a commercial enterprise. But distrust seeped in during the early stages and resulted in the launch of three separate businesses. One company was formed by Emmanuelle Charpentier and partners: CRISPR Therapeutics, initially based in Switzerland but now with headquarters also in Cambridge, Massachusetts. A second company is Editas Medicine, formed around Feng Zhang and George Church. Jennifer Doudna was also involved initially in establishing Editas, but left abruptly when she learned that Zhang and the Broad Institute, without alerting her, had filed for a CRISPR-editing patent in apparent opposition to the application Doudna and Charpentier had submitted together with their respective institutions seven months earlier. Doudna instead collaborated with several other CRISPR pioneers, including Rodolphe Barrangou, and one of her own former students to form Intellia Therapeutics. The patent priority disputes between Doudna-Charpentier and Zhang/Broad Institute have continued for eight years, with an interim solution approving some patent claims for each party, but court cases continuing to this day.
III. the types of crispr editing
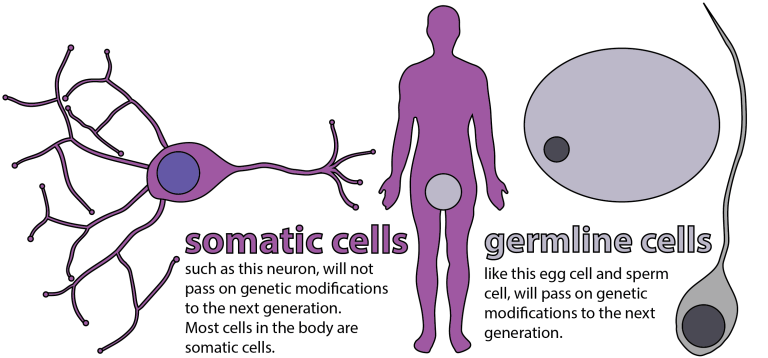
One can distinguish three types of CRISPR editing, differing in the inheritability of the engineered changes to DNA. Figure 4 indicates the distinction between two of these types, somatic cell and germline cell editing. The vast majority of human cells are somatic cells uninvolved in reproduction. Hence, edits made to DNA within somatic cells are not passed along to the next generation, though they can be critical for treating diseases centered in those cells and related to gene mutations. A prominent example is sickle-cell anemia, which involves a shape distortion of red blood cells caused by a kink in hemoglobin protein arising from a single-base mutation that occurs in both copies of a single gene in the two chromosomes contributed by a person’s biological father and mother.
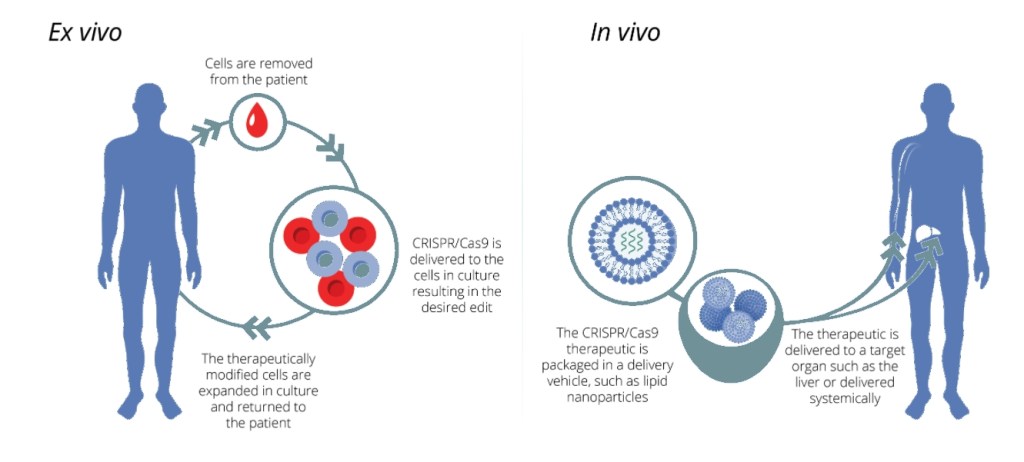
The treatment of sickle-cell anemia is best done ex-vivo (see Fig. 5) by extracting blood stem cells, editing them outside the body, and then reinjecting the modified cells where they can produce healthy hemoglobin. Other somatic edits can be done in-vivo (Fig. 5) if the targeted cells are not easily removed from the body and there is an appropriate delivery mechanism to get the CRISPR complex to the nucleus of the particular cells of interest inside the body.
Somatic cell edits are not inheritable. In contrast, germline edits made to cells involved in reproduction – egg or sperm cells, or embryos – affect every cell in offspring and are inheritable, and thus capable of eliminating a genetic disease from all descendants, as long as future mates do not themselves carry the genetic disease. Germline editing is quite controversial. The potential to eliminate a disease-causing genetic defect or susceptibility from an entire line of descendants is a powerful motivator for developing it. But it could also be used, in principle, not for such therapeutic purposes but rather to introduce desired “enhancements,” such as increased strength or intelligence, into “CRISPR babies” and a line of descendants, or even into an entire “caste” of genetically enhanced humans, such as “super soldiers” (about which we will say more in part II of this post). Among such genetic enhancements might be edits to produce a line of drought-resistant crops or stronger and healthier livestock.
At the moment, however, CRISPR editing is not yet sufficiently precise to eliminate the possibility of serious side-effects arising, for example, from unplanned off-target DNA modifications. Until such problems (to be discussed further in the following section) are understood and mitigated, it may be quite dangerous to introduce germline edits even for therapeutic purposes. One may be substituting one or more unforeseen genetic disorders in place of the one to be mitigated or enhanced.
It thus came as a shock to the CRISPR research community, and indeed to the entire world, when Chinese research-entrepreneur He Jiankui announced in November 2018 that he had successfully produced the first germline-edited CRISPR babies. Jiankui had recruited volunteer Chinese couples involving a husband who had had AIDS or tested positive for the HIV virus and an HIV-negative wife. For two such couples, Jiankui removed the HIV virus from sperm cells donated by the father and then edited egg cells from the mother after they had been fertilized in vitro by the cleaned sperm. The CRISPR-Cas9 edit targeted the CCR5 gene that encodes a protein which serves as a receptor for the HIV virus.
He Jiankui’s stated goal was to produce offspring who would be unsusceptible to later HIV infection. The edited, fertilized eggs produced embryos that were then reimplanted in the mother, and in one case this procedure resulted in the birth of healthy twin girls. It turned out that in one of the twins the edits had affected only one of her two chromosomes, so that her body would still be capable of producing the CCR5 protein, and in both there were also undesired off-target edits and some unedited cells. These results led one U.S. researcher to comment “The first attempt to hack the code of life and, ostensibly, improve the health of human babies had in fact been a hack job.”
Chinese researchers had, for a few years prior to Jiankui’s announcement, already carried out tests of germline CRISPR editing, but only on non-viable human embryos. Although He Jiankui had apparently anticipated being declared a national hero for his groundbreaking work, it violated a Chinese regulation against germline genome editing in humans. Jiankui was prosecuted for this violation in a Chinese court, which declared “In order to pursue fame and profit, [he] deliberately violated the relevant national regulations and crossed the bottom lines of scientific and medical ethics.” He Jiankui was sentenced in December 2019 to three years in prison, fined $430,000 and banned for life from working in reproductive science.
In the wake of He Jiankui’s revelation of his creating the first CRISPR babies, bioscientists worldwide have gotten together to accelerate earlier discussions about ethical rules for germline editing. An international commission formed from various national academies of science after the He Jiankui announcement produced a report in 2020 that did not call for a moratorium on germline editing research, as some had promoted, but rather said that such editing “might in the future provide a reproductive option” for couples who have, or carry genes favoring, genetic diseases. The report noted the present dangers involved in germline editing, and that such editing even for therapeutic purposes was often not medically necessary, i.e., other treatment options exist. In the end, the report supported “defining a responsible pathway for [future] clinical use of heritable human genome editing” when it is deemed to be both safe and medically necessary. This appears at the moment to represent the consensus among the world’s scientists who work on genome editing.
Germline editing is intended to affect a full line of descendants from a single reproduction. In contrast, the third type of genomic modification, gene drives, use CRISPR-Cas9 editing to introduce engineered genes throughout an entire population of rapidly reproducing species. As we have described in a previous post, they do this by inserting an externally generated CRISPR complex, including a desired-trait payload gene, on a small DNA loop (called a plasmid) into the target cells of a small sample of the population. When the Cas9 scissors enzyme cleaves the targeted DNA sequence in one chromosome, the entire CRISPR-Cas9-payload gene complex is copied in its place via homology-directed repair. Because the CRISPR-Cas9-payload gene complex is thus introduced within one chromosome of the targeted individuals, it automatically cuts and replaces the similar gene in the matched chromosome supplied by the other parent.
Once the edit is thus introduced in both chromosomes, it makes inheritance of the engineered trait nearly inevitable when the modified individuals mate with unmodified members of the wild population, because the entire CRISPR complex in a chromosome contributed by the modified parent will once again copy itself into its paired chromosome from the other parent. Since a number of generations are needed to spread the engineered trait through the entire wild population, gene drives are most effective for rapidly reproducing species, such as mosquitoes, where the edit can reduce the probability that the population act as vectors carrying diseases such as malaria. While the goals may be noble, it is critical to understand the downsides of applying such new technology; a previous attempt to eradicate pests with DDT had serious negative impacts on humans, as emphasized by Rachel Carson in her book Silent Spring and in our previous post on scientific tipping points. Gene drives carry their own ethical concerns about ecosystem disruption, which we have discussed in our earlier post.
IV. current crispr research and clinical forefronts
Following the 2013 demonstration that CRISPR-Cas9 editing could be done in the nucleus-containing cells of humans and other animals, there has been an explosion (see Fig. 6) in the number of research efforts and clinical trials exploring CRISPR editing applications. We briefly review some of those efforts in this section.

Clinical Trials:
The therapeutic companies established by the CRISPR pioneers, together with a number of university groups, hospitals, pharmaceutical companies and research institutes in both the U.S. and China, are currently leading a significant number of ongoing clinical trials of CRISPR-based therapies. Among the single-gene disorders, sickle-cell anemia “is one of the best candidates for ex vivo editing because it involves blood cells that can be easily extracted and returned…The simplicity of the [single-base] genetic glitch and the severity of the syndrome make it a perfect candidate for gene editing.” The first therapeutic treatment of humans in the U.S. via CRISPR-Cas9 editing was done in 2019, as part of a clinical trial, on an African-American woman with sickle cell anemia. In addition to sickle-cell clinical trials, others have been initiated to test gene editing to treat the blood disorder thalassemia and Leber congenital amaurosis, a common cause of childhood blindness, caused by a mutation in the gene that makes light-receptor cells in the eyes. Here the treatment is in vivo, because eye cells cannot be extracted and returned like blood cells. A tailored virus has to be used as a delivery vehicle to transport the CRISPR-Cas9 into the targeted cells beneath the retina.
A number of clinical trials launched in China are focused on testing CRISPR-based cancer immunotherapies. Cancerous cells often protect themselves from the body’s immune system by triggering the expression of a gene that produces a protein called PD-1, which halts the cell’s immune response. CRISPR-Cas9 can be used to disable that gene ex vivo in T-cells extracted from the patient’s blood. The T-cells are white blood cells involved in the immune response. An analogous trial led by researchers at the University of Pennsylvania is testing edits that not only knock out the PD-1 gene, but additionally insert a gene that targets the patients’ tumors. The trials so far have demonstrated at least that the edited T-cells can be tolerated and thrive once returned to the patient’s blood stream.
So far, clinical trials have tested, or at least been launched to test, this cancer immunotherapy approach on patients with cancer of the lungs, esophagus, bladder, prostate and kidney, among others. In addition, according to Isaacson, “Mammoth Biosciences, a company that Doudna founded with two of her graduate students, is designing diagnostic tools based on CRISPR that can be used on tumors to identify quickly and easily the DNA sequences associated with different types of cancers. Then precision treatments can be tailored for each patient.”
Ongoing research not yet at the level of clinical trials is also developing an alternative approach to managing cancer. It involves CRISPR editing to add an extra P53 gene to humans. This gene encodes a protein that suppresses the growth of cancerous tumors, responding to damaged DNA in such a way as to prevent cancerous cells from dividing. Cancers proliferate when humans’ one copy of this gene is damaged, or goes unexpressed. Other animals naturally have more copies of the P53 gene – for example, elephants have twenty copies and almost never get cancer.
Other clinical trials have begun to test treatments “for angioedema (a hereditary disease that causes severe swelling), acute myeloid leukemia, super-high cholesterol, and male pattern baldness.”
Improving CRISPR-Cas9:
Gene editing in humans is not without potentially serious side-effects. Guide RNAs “tend to have relatively high mismatch tolerance; hence, Cas9 commonly cleaves off-target sites that have sequences similar to those of target genes.” In a 2018 article in the journal Nature Biotechnology, researchers from the Wellcome Sanger Institute in the UK carried out a full systematic study of the entire genome in both mouse and human cells that had been subjected to CRISPR-Cas9 editing. While previous studies had revealed very limited “unforeseen mutations from CRISPR-Cas9 in the DNA at the genome-editing target site,” the authors found that the editing “deleted thousands of DNA bases – some of them that were not close to the spot of the initial edit…some of the deletions can silence genes that are supposed to be active…and activate some genes that should be turned off – particularly genes that can cause cancer.” Furthermore, the authors noted that some of “the genetic damage can be difficult to discover through standard DNA tests.”
Another problem arises because Cas9 sometimes triggers human anti-Cas9 immune response, which “causes low editing efficiency and could lead to a serious immune storm in patients receiving CRISPR-Cas9 treatment.” So researchers are working to design improved, more accurate and less error-prone, versions of both the guide RNA and the Cas9 enzyme itself. For example, one modification to Cas9 allows it to recognize a wider range of the special, so-called PAM (protospacer adjacent modif), sequences that tell Cas9 where to cut, thereby making the editing more versatile.
Turning CRISPR Off:
Perhaps a more worrisome problem than CRISPR side-effects is its potential misuse. Just as Robert Oppenheimer, who had led the Manhattan Project to develop the atomic bomb, came to fear that the monster he had helped to create might be used, along with even more powerful hydrogen bombs, to destroy the world, Jennifer Doudna, who had demonstrated the capabilities of CRISPR-Cas9 for rapid gene-editing, came to fear the potential misuse of the technology that had so much promise for good. In 2014 she had a nightmare in which a prominent researcher asked her to meet someone who wanted to learn about gene editing. As Walter Isaacson tells the story in his book The Codebreaker, “Sitting in front of her, with pen and paper ready to take notes, was Adolf Hitler with the face of a pig. ‘I want to understand the uses and implications of this amazing technology you’ve developed,’ he said.”
Doudna’s concerns were furthermore excited by a conference talk describing “how a virus could be engineered to carry CRISPR components into mice and edit a gene so that the mice would get lung cancer.” Doudna herself wrote in her own book A Crack in Creation: “Emmanuelle and I, and our collaborators, had imagined that CRISPR technology could save lives by helping to cure genetic disease. Yet as I thought about it now, I could scarcely begin to conceive of all of the ways in which our hard work might be perverted…Could I and other concerned scientists save CRISPR from itself … before a cataclysm occurred?”
Doudna raised public awareness of her concerns. In 2016 the Worldwide Threat Assessment issued annually by the U.S. Director of National Intelligence for the first time discussed “genome editing” as a potential weapon of mass destruction. This led directly to DARPA’s 2016 launch of its Safe Genes program of research funding “to mitigate the risks and security concerns related to the accidental or intentional misuse of such technologies and, at the same time, enable the pursuit of novel genetic solutions that support public health and military force protection and readiness.” Doudna herself received one of the first of the Safe Genes grants, with the goal of creating a CRISPR “kill switch” that “might someday be capable of disabling weapons employing CRISPR.”
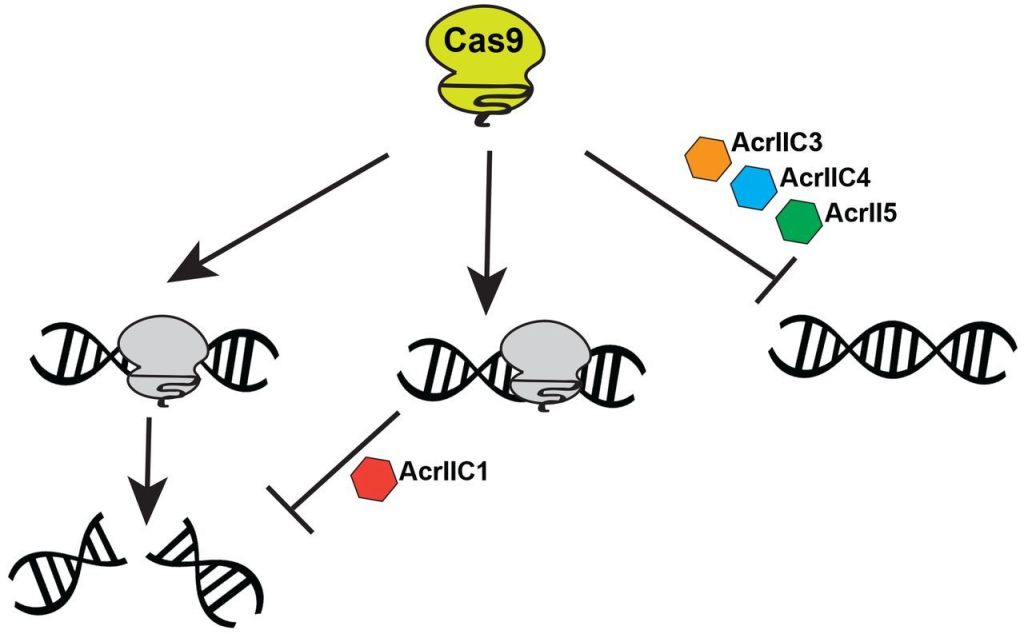
Nature and evolution had already provided a solution. Just as CRISPR itself was discovered initially in the immune system many bacterial species had evolved over eons to destroy invading viruses that had been encountered in preceding generations, the CRISPR kill switch, or anti-CRISPR was discovered accidentally, in the form of proteins produced by some genes in viruses whose genome had evolved to overcome the bacterial immunity. These were initially discovered by a group at the University of California at San Francisco. By now more than 50 anti-CRISPR proteins have been characterized, each with its means of blocking the cut-and-paste action of CRISPR systems: some inhibit the splicing action of the Cas9 enzyme, while others inhibit the ability of the CRISPR complex to lock onto its targeted DNA segment – see Fig 7.
Doudna’s team first demonstrated that anti-CRISPRs introduced into human cells could halt Cas9 gene-editing activity in its tracks and limit off-target activity as well. As Isaacson summarizes: “The anti-CRISPRs could be engineered to regulate gene-editing systems. That would be useful for medical applications that needed to time-limit a CRISPR edit, and they could be used as a defense against systems created by terrorists or malevolent enemies. Anti-CRISPRs could also be used to shut off gene drives…”
A German team has furthermore demonstrated that by combining CRISPR-Cas9 and anti-CRISPR they can create a selective DNA editor that works in liver cells, but not in neurons or muscle. Yet recognition of the potential applications and proof-of-principle laboratory experiments on isolated cells are not yet the equivalent of an executable therapy. Doudna has recently asked: “How do you actually use these in a way that will provide meaningful control? That’s certainly where that whole anti-CRISPR field needs to go. It just hasn’t gone there yet.”
If anti-CRISPR proteins provide a useful kill switch for CRISPR-based DNA editing, another DARPA-funded project has very recently resulted in an extremely promising CRISPR-based “on-off” switch (see Fig. 8) for gene expression, i.e., for the actual assembly of targeted proteins under instructions encoded within specific genes. In its more conventional combination with Cas9 enzymes, CRISPR cuts double-stranded DNA, leaving the repairs to “in-house” cellular repair mechanisms, sometimes influenced by the presence of payload DNA templates contained in the CRISPR-Cas9 complex itself. These repair mechanisms may induce unintended changes as well as the intended one, and in any case, the changes to DNA are permanent.

The new approach does not make permanent alterations to the DNA itself, but rather adds chemical tags to a gene targeted by CRISPR – a process known as methylation – which inhibit the expression of the gene. The research team refers to the complex that accomplishes this task as CRISPRoff. But in addition they developed a CRISPRon complex, which employs an enzyme that removes the methyl groups attached to DNA segments targeted by CRISPRoff. This gives the on-off switch illustrated in Fig. 8. As MIT Professor Jonathan Weissman, who led the team, summarizes: “The big story here is we now have a simple tool that can silence the vast majority of genes. We can do this for multiple genes at the same time without any DNA damage, with great deal of homogeneity, and in a way that can be reversed. It’s a great tool for controlling gene expression.”
The team demonstrated the operation of the switch within stem cells by successfully turning down the expression of the Tau protein, which is implicated in Alzheimer’s disease – a very promising result, though not yet accompanied by a delivery mechanism to introduce CRISPRoff selectively to appropriate cells within a human adult. It will also be very useful in further elucidating the role played by individual genes (whose expression can be turned off or on) in various phenotype features and diseases of humans.
Other CRISPR-associated Enzymes:
Cas9 is only one of several CRISPR-associated enzymes found in bacterial genomes. While Cas9 makes one double-stranded set of cuts in DNA, two other enzymes – Cas12a and Cas13 – have come to prominence in the rapid developments to exploit CRISPR technology to help deal with the COVID-19 pandemic. Cas12 performs a double-stranded cleavage, like Cas9, but then continues to chop up any nearby single-stranded molecules. If the CRISPR-Cas12 system is supplemented by a diagnostic molecule, the chopping of that molecule could cause a glowing signal of the presence of an invading DNA-based virus to which the CRISPR guide RNA is matched. Two of Doudna’s students showed, for example, that CRISPR-Cas12 could detect human papillomavirus (HPV), a sexually transmitted infection.
On the other hand, the SARS-COV-2 virus that causes COVID-19 is an RNA-based virus and is thus not directly affected by Cas9 or Cas12. However, Feng Zhang’s team at the Broad Institute showed that Cas13 is a similar enzyme to Cas12, but performs only single-strand cuts on RNA, before going into a similar cutting frenzy to Cas12. Both Cas12 and Cas13 have now been adapted to produce very promising, quick and accurate, COVID-19 diagnostic tests, marketed by companies Mammoth Biosciences (associated with Doudna’s group) and Sherlock Biosciences (associated with Zhang), both expected to come to market shortly. DARPA is furthermore funding research into the possibility of using CRISPR-Cas13 as a treatment for COVID or other RNA-viruses, one which would target invading viruses matched to the engineered guide RNA and chop them to bits, much as bacteria have evolved to do over billions of years.
Meanwhile, a new miniature protein CasMINI has been engineered by Stanley Qi and collaborators at Stanford University, with function similar to that of Cas12a, particularly for gene activation. The reduced length of the CasMINI molecule (~40% of the length of Cas9 or Cas12a) allows it more easily to be attached as a payload to viruses used to transport CRISPR complexes across human cell boundaries for in vivo treatments, expanding the range of possible clinical applications. There is also hope that CasMINI may be less susceptible to triggering human immune reactions than Cas9 or Cas12a.
— To be continued in Part II —
